Detection of Inducible Clindamycin Resistance among Staphylococcal Isolates from Different Clinical Specimens in Northwestern Nigeria
Kumurya A. S.*
Department of Medical Laboratory Science, Faculty of Allied Health Sciences, Bayero University, Kano, Nigeria
Abstract
Background: Macrolide (MLSB) resistance is the most widespread and clinically important mechanism of resistance encountered with Gram-positive organisms. Resistance may be constitutive (cMLSB phenotype) or inducible (iMLSB phenotype). The iMLSB phenotypes are not differentiated by using standard susceptibility test methods, but can be distinguished by erythromycin-clindamycin disk approximation test (D-test) and demonstration of resistance genes by molecular methods. Aims: To demonstrate in vitro iMLSB in erythromycin-resistant (ER) and clindamycin-susceptible (CLI-S) clinical isolates of Staphylococcus aureus and interpretation of susceptibility tests to guide therapy. Materials and Methods: One hundred isolates of Staphylococcus aureus were recovered from various clinical specimens. All the Staphylococcus aureus were identified by conventional microbiological methods including colony morphology, Gram stain, catalase, slide coagulase and tube coagulase. Polymerase chain reaction (PCR) was used to amplify both the S. aureus specific sequence gene and mecA gene of 100 isolates with the amplicon size of 107 and 532 bp respectively. Antibiotic susceptibility testing was performed by Kirby Bauer disc diffusion method. Erythromycin-resistant isolates were examined for iMLSB by using double disk approximation test (D-test) at 15 mm disk separation. Results: All the isolates (n=100) expressed S. aureus specific sequence gene in their PCR products. Only 5 isolates (5.0%) were confirmed as Methicillin resistant Staphylococcus aureus (MRSA) based on the detection of mecA gene. Seven percent of Staphylococcus aureus were ER among which 4.0% were Methicillin susceptible Staphylococcus aureus (MSSA) and 3.0% were MRSA. The inducible MLSB (iMLSB) phenotype was detected in 3 MRSA isolates and all the 4 erythromycin-resistant MSSA strains were positive for the cMLSB phenotypes. Sixty per cent of iMLSB phenotypes were observed to be methicillin-resistant. All the isolates (MSSA and MRSA) were susceptible to vancomycin. Conclusions: The identification of multi-resistant MSSA with in vitro inducible clindamycin resistance at our health institutions raises concern of clindamycin treatment failures with methicillin-resistant infections. It is recommended that microbiology laboratories should include the D-test for inducible resistance to clindamycin in the routine antibiotic susceptibility testing.
Keywords
D-Test, Inducible Clindamycin Resistance, Staphylococcus aureus
Received: April 9, 2015
Accepted: April 22, 2015
Published online: May 27, 2015
@ 2015 The Authors. Published by American Institute of Science. This Open Access article is under the CC BY-NC license. http://creativecommons.org/licenses/by-nc/4.0/
1. Introduction
Infections due to staphylococci, especially methicillin resistant S. aureus (MRSA) are increasing among in-patient clinics and treatment of these infections pose difficulties (Manian et al., 2003; Maltezou et al., 2006). Macrolides, lincosamides and streptogramin (MLS) antibiotics are used in the treatment of staphylococcal infections. Macrolide (MLSB) resistance is the most common and clinically important mechanism of resistance encountered with Gram-positive organisms. MLS antibiotics are structurally unrelated, but microbiologically related antibiotics because of their similar mode of action. Resistance may be constitutive (cMLSB phenotype) or inducible (iMLSB phenotype). The iMLSB phenotypes are not differentiated by using standard susceptibility test methods, but can be distinguished by erythromycin-clindamycin disc approximation test (D-test) and demonstration of resistance genes by molecular methods (Sesli et al., 2008; Colakoğlu, 2008; Dinç et al., 2009; Pal et al., 2010; Gatermann et al., 2007). Resistance to MLS antibiotics occurs either through target site modification, efflux of antibiotics or drug modification (Leclercq, 2002). MLS antibiotics inhibit bacterial protein synthesis by binding to 23srRNA, which is a part of the large ribosomal subunit. In target site modification methylation of the A2058 residue, located in the conserved domain V of 23srRNA. The development of resistance in Staphylococcus species to macrolide, lincosamide and streptogramin B has limited the use of these antibiotics. Macrolide resistance may be due to enzymes encoded by a variety of erm genes-MLSB phenotype and may be constitutive (cMLSB phenotype) or inducible (iMLSB phenotype). Another mechanism is active efflux pump encoded by the mrsA gene (MS phenotype). The MS and iMLSB phenotypes are indistinguishable by using standard susceptibility test methods, but can be distinguished by erythromycin-clindamycin disk approximation test (D-test) and demonstration of resistance genes by molecular methods (Drinkovic et al., 2001 and Steward et al., 2005). The aim of this study was to determine the rate of inducible clindamycin resistance in both methicillin-resistant and susceptible strains of Staphylococcus in our hospital as data describing iMLSB prevalence among Staphylococcus isolates in this region is unknown.
2. Materials and Methods
A total of 100 consecutive non – duplicated Staphylococcus aureus isolates were obtained from clinical samples in 8 health institutions (Microbiology department) across Northwestern Nigeria. The isolates were collected for duration of ten months from July 2009 to April 2010. The quality control and rejection criteria of specimen were followed (Muhammad et al., 2010). All plates were examined for Staphylococcus by colonial morphology on nutrient agar (Cheesbrough, 2000). Catalase, coagulase, DNase tests and test for heamolysin were performed on all the isolates. Staphylococcus aureus ATCC 25923 was used as a reference control organism. All confirmed Staphylococcus aureus isolates were stored in 16% v/v glycerol broth at -80oC.
Antibiotic susceptibility testing was performed by Kirby Bauer disc diffusion method according to the National Committee for Clinical Laboratory Standards (now Clinical Laboratory Standards institute) guidelines (NCCLS, 2006). The antibiotics (Mast Diagnostics, UK) tested includes penicillin (10µg), fusidic acid (10µg), ciprofloxacin (5µg), gentamycin (10µg), erythromycin (15µg), chloramphenicol (30µg), tetracycline (30µg), trimethoprin (2.5µg), augumentin (30µg), ofloxacin (10µg), peflacine (5µg), streptomycin (30µg), cefuroxime (30µg), cotrimoxole (25µg) and clindamycin (2µg). Staphylococcus aureus (ATCC25923) was the control strain in every test run. Methicillin resistance was detected by Polymerase chain reaction (PCR). Amplification of both the S. aureus specific sequence gene and mecA gene of 100 isolates with the amplicon size of 107 and 532 bp was done by PCR test. One MSSA control strain (ATCC 25923) was included in each batch of PCR run. (Wielders et al., 2001; Meshref and Omer, 2011).
Disk approximation testing (D-test) was performed for each isolate according to Clinical and Laboratory Standards Institute (CLSI) method (CLSI, 2005). A 0.5 McFarland suspension was prepared in normal saline for each isolate and inoculated on Mueller-Hinton agar plate. Clindamycin (CLI)-2 μg and erythromycin (ER)-15μg disks were placed 15 mm apart edge to edge manually. Plates were incubated at 35 0 C for 24 h and zone diameters were recorded. Induction test categories were interpreted (Steward at al., 2005).
3. Results
A total of 100 Staphylococcus aureus isolates were included, of which 5.0% were MRSA and 95.0% MSSA. Categorization of the isolates along with sources is depicted in (Table 1).
Table 1. Sources and categorization of Staphylococcus aureus isolates
| Specimen | MRSA | MSSA | TOTAL |
| Wound swabs | 3(9.7) | 28 (90.3) | 31 (31.0) |
| Ear swabs | 0(0.0) | 9 (100) | 9 (9.0) |
| Blood culture | 0(0.0) | 8 (100) | 8 (8.0) |
| Urine | 2(7.7) | 24 (92.3) | 26 (26.0) |
| High vaginal swabs | 12 (100) | 12(100) | 12 (12.0) |
| Sputum | 0(0.0) | 11 (11.0) | 11 (11.0) |
| Semen | 0(0.0) | 1 (100) | 1 (100) |
| Urethral swabs | 0(0.0) | 2 (100) | 2 (100) |
| Total | 5 (5.0) | 95 (95.0) | 100 (100) |
Seven (7.0%) clinical isolates which showed erythromycin resistance were tested for inducible resistance by double disk approximation test. Out of 7 erythromycin-resistant strains 3 (3.0%) were iMLSB phenotypes [60.0% MRSA (3/5)]. All the iMLSB phenotypes were observed to be methicillin-resistant isolates. D-test yielded two distinct induction phenotypes, D-zone phenotype [Figure 1] was observed in 78 (18.13%) and D + phenotype [Figure 2] in 22 (5.34%) isolates. Both D and D + results were considered positive for CLI induction- iMLSB phenotypes. One hundred and twenty-seven (29.53%) isolates showed ER resistant and CLI-susceptible zone diameters with no blunting of the zones (MS phenotype). Two hundred and two (46.97%) isolates showed ER and CLI resistance (cMLSB phenotype), of which 98 were MRCNS, 85 MRSA and the rest were methicillin-sensitive [Table 2]. No hazy D zone (HD) phenotype was observed.
Susceptibility of iMLSB phenotypes isolated were ampicillin 37.5%, amoxiclav 39.13%, cefepime 12.5%, ceftriaxone 50.0%, cephotaxime 62.5%, cefoperazone-sulbactam 60.86%, cephalexin 23.07%, ciprofloxacin 78.78%, doxycycline 69.56%, linezolid 100%, netilmicin 46.60%, piperacillin-tazobactum 69.56% and vancomycin 100%.
Table 2. MLSB resistance phenotypes in S. aureus isolates
| Strains | MLSB phenotypes (%) | |
| Constitutive MLSB Phenotypes (%) | Inducible MLSB phenotypes (%) | |
| MSSA | 4 (100) | 0 (0) |
| MRSA | 0 (0) | 84 (100) |
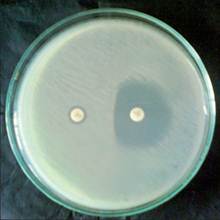
Figure 1. Double disc test (D test) showing flattening of the zone of inhibition around clindamycin disc proximal to erythromycin disc (D shaped zone of inhibition)
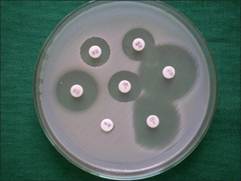
Figure 2. Inducible MLSB isolate- D phenotype
4. Discussion
In our study, 851 Staphyloccocal isolates were obtained over a period of nine months (2008-09) in which 50.52% were ER-resistant. Among the ER-resistant S. aureusiMLSB resistance was observed in 24.63% (51/207) similar to that reported by Fiebelkornet al., 2003; Jorgensen et al., 2004 and Gadepalli et al., 2006. Some investigators have reported a higher incidence of iMLSBresistance(Goyalet al., 2004; Merino et al., 2005; O'Sullivanet al.,2006; Patel et al., 2006; Angel et al.,2008; Cirajet al., 2009) while others have indicated lower incidence. (Jenssen et al., 1987; Hamilton-Miller and Shah, 2000; Delialioglu et al., 2005; Huanget al., 2006; Rahbar andHajia, 2007). We observed almost a similar rate of iMLSB resistance among S. aureus (24.63%) and CNS (22.42%), while others have reported variable results. The different patterns of resistance observed in various studies are because iMLSB resistance varies by geographical region, age group, methicillin susceptibility and even from hospital to hospital. As observed in studies by Steward et al., and Schreckenberger et al., our CLI induction results also showed two phenotypes, D (18.13%) and D + (5.34%) phenotypes and both are considered to be positive D-zone test (Steward et al., 2001 ad Schreckenberger et al., 2004). All the iMLSB isolates were susceptible to vancomycin while 78.78% to ciprofloxacin. Rahbar and Hajia also found all iMLSB isolates susceptible to vancomycin (Rahbar and Hajia, 2007).
Due to the emergence of resistance to antimicrobial agents accurate drug susceptibility data of the infecting microbe is an essential factor in making appropriate therapeutic decisions. MLSB resistance is the most widespread and clinically important mechanism of resistance encountered with Gram-positive organisms due to the production of methylases and efflux proteins. In vitro susceptibility testing for clindamycin may indicate false susceptibility by the broth microdilution method and by disk diffusion testing with erythromycin and clindamycin disks in nonadjacent positions. Erythromycin-clindamycin disc approximation test or D-test is a simple, reliable method to detect inducible resistance to clindamycin in erythromycin-resistant isolates of Staphylococci (Steward et al., 2001 and Fiebelkorn et al., 2003). Sensitivity of D-test performed at 15-20 mm disk spacing was 100% when correlated with detection of erm and msr genes by polymerase chain reaction (PCR) (Steward et al., 2001; Fiebelkorn et al., 2003; Jorgensen et al., 2004; Merino et al., 2005 and O'Sullivan et al., 2006).
Clinically, bacterial strains exhibiting iMLSB have a high rate of spontaneous mutation to constitutive resistance and use of non-inducer antibiotics such as clindamycin can lead to selection of constitutive mutants at frequencies of 10 -7cfu (McGehee et al., 1968; Goyal et al., 2004; Schreckenberger et al., 2004;O'Sullivan et al., 2006). McGehee and other investigators have confirmed this rapid in vitro conversion of inducible to constitutive MLS B resistance in Staphylococci (McGehee et al., 1968 and Panagea et al., 1999; O'Sullivan et al., 2006). There have also been a number of reported clindamycin or lincomycin therapy failures in serious infections due to Staphylococci with iMLSB resistance, indicating that it is not uncommon (Drinkovic et al., 2001 and Panagea et al., 1999). Clindamycin has long been an attractive option in the treatment of skin and soft tissue infections (SSTI) and serious infections because of its efficacy against MRSA and MSSA, as well as anaerobes. This has led to questioning the efficacy of clindamycin use against any erythromycin-resistant Staphylococci spp. However, if inducible resistance can be reliably detected on a routine basis in clinically significant isolates, clindamycin can be safely and effectively used in patients with true clindamycin-susceptible strains.
In the present study, 29.53% of erythromycin-resistant Staphylococcal isolates showed true clindamycin susceptibility (MS phenotype). Patients with infections caused by such isolates can be treated with clindamycin without emergence of resistance during therapy.
The high frequency of methicillin-resistance isolates (87.12%) with in vitro inducible clindamycin resistance at our institute raises concern of clindamycin treatment failures with methicillin-resistant infections.
5. Conclusion
It can be concluded that it is important for laboratories to be aware of the local prevalence of iMLSB isolates. On the basis of their data they can choose whether or not to perform the D-test routinely. The D-test is an easy, sensitive, and reliable means for detection of iMLSB strains in a clinical laboratory setting without specialized testing facilities. The prevalence of iMLSB may change over time with the emergence of strains with different sensitivity patterns, so periodic surveys should be performed if testing is not routine.
Recommendations
Currently, some authors recommend avoidance of clindamycin for treatment of complicated infections which may have a high bacterial burden, such as abscesses or osteomyelitis. Conversely labelling all erythromycin-resistant staphylococci as clindamycin resistant would prevent the use of clindamycin in infections caused by truly clindamycin susceptible isolates. Clindamycin may be useful for non-MLSBi infections especially less severe S. aureus infections. This study reflects the prevalence of MLSBi at a tertiary care centre; however prevalence may differ from institute to institute. Microbiological laboratories should adopt testing for MLSBi among S. aureus isolates and report isolates exhibiting MLSBi as clindamycin resistant.
Acknowledgements
I thank the management of the health institutions for their ethical permission to collect Staphylococcus aureus isolates from their facilities. My appreciation also goes to the entire staff of Medical Microbiology Laboratories of the health institutions for their valuable contributions and assistance in the collection of the isolates.
References
- Angel MR, Balaji V, Prakash J, BrahmadathanKN, Mathews MS (2008). Prevalence of inducible clindamycin resistance in gram positive organisms in a tertiary care centre. Indian J Med Microbiol 26:262-4.
- Ciraj AM, Vinod P, Sreejith G, Rajani K (2009). Inducible clindamycin resistance among clinical isolates of staphylococci.Indian J PatholMicrobiol52:49-51.
- Clinical and Laboratory Standards Institute (CLSI) (2005). Performance Standards for Antimicrobial disk susceptibility Tests: Approved Standard M2-A7. 11th ed. Clinical and Laboratory Standards Institute Wayne, Pa.
- Corinne JH, Henry FC (1989). Methicillin-resistant staphylococci: Detection methods and treatment of infections. Antimicrob Agents Chemother33:995-9.
- Delialioglu N, Aslan G, Ozturk C, Baki V, Sen S, Emekdas G (2005). Inducible clindamycin resistance in staphylococci isolated from clinical samples. Jpn J Infect Dis58:104-6.
- Drinkovic D, Fuller ER, Shore KP, Holland DJ, Rod Ellis Pegler (2001). Clindamycin treatment of Staphylococcus aureus expressing inducible clindamycin resistance. J AntimicrobChemother48:315-6.
- Fiebelkorn KR, Crawford SA, McElmeel ML, Jorgensen JH (2003). Practical disk diffusion method for detection of inducible clindamycin resistance in Staphylococcus aureus and coagulase-negative staphylococci. J ClinMicrobiol41:4740-4.
- Gadepalli R, Dhawan B, Mohanty S, Kapil A, Das BK, Chaudhry R (2006). Inducible clindamycin resistance in clinical isolates of Staphylococcus aureus. Indian J Med Res123:571-3.
- Steward CD, Raney PM, Morrell AK, Williams PP, McDougal LK, Jevitt L (2005). Testing for induction of clindamycin resistance in erythromycin-resistant isolates ofStaphylococcus aureus. J ClinMicrobiol43:1716-21.
- Jorgensen JH, Crawford SA, McElmeelMLandFiebelkorn KR (2004). Detection of inducible clindamycin resistance of Staphylococci in conjunction with performance of automated broth susceptibility testing. J ClinMicrobiol42:1800-2.
- Goyal R, Singh NP, Manchanda V, Mathur M (2004). Detection of clindamycin susceptibility in macrolides resistant phenotypes of Staphylococcus aureus. Indian J Med Microbiol22:251-4.
- Hamilton-Miller JM, Shah S (2000). Patterns of phenotypic resistance to the macrolide-lincosamide-ketolide-streptogramin group of antibiotics in staphylococci. J AntimicrobChemother46:941-9.
- Huang H, Flynn NM, King JH,Monchaud C, Morita M, Cohen SH (2006). Comparisons of community-associated methicillin-resistant Staphylococcus aureus (MRSA) and hospital-associated MSRA infections in sacramento, California. J ClinMicrobiol44:2423-7.
- JenssenWD, Thakker-Varia S, Dubin DT, Weinstein MP (1987). Prevalence of macrolides-lincosamides-streptogramin B resistance and erm gene classes among clinical strains of staphylococci and streptococci. Antimicrob Agents Chemother31:883-8.
- Meshref, A. A. and Omer, M. K.(2011). Detection of (mecA) gene in methicillin resistantStaphylococcus aureus(MRSA) at Prince A / RhmanSidery Hospital, Al-Jouf, SaudiArabia.Journal of Medical Genetics and Genomics.3 (3): 41 – 45.
- Merino L, Torres MJ, Cantos A, Ruiz M, Aznar J (2005). Detection of inducible clindamycin resistance in cutaneous Staphylococci clinical isolates by phenotypic and genotypic method. Abstract number: 1134_02_427.15 thEuropean Congress of Clinical Microbiology and Infectious Diseases Copenhagen / Denmark, April 2-5.
- O'Sullivan MV, Cai Y, Kong F, Zeng X, Gilbert GL (2006). The influence of disk separation distance on the accuracy of the disk approximation testing for inducible clindamycin resistance in Staphylococcus spp. J ClinMicrobiol44:4072-6.
- Panagea S, Perry JD, Gould FK (1999). Should clindamycin be used in treatment of patients with infections caused by erythromycin-resistant staphylococci? J AntimicrobChemother44:581-2.
- Patel M, Waites KB, Moser SA, Cloud GA, Hoesley CJ (2006). Prevalence of inducible clindamycin resistance among community and hospital-associated Staphylococcus aureus isolates. J ClinMicrobiol44:2481-4.
- Rahbar M and Hajia M (2007). Inducible clindamycin resistance in Staphylococcus aureus: A cross-sectional report. Pak J BiolSci10:189-92.
- Schreckenberger PC, Ilendo E, Ristow KL (2004). Incidence of constitutive and inducible clindamycin resistance in Staphylococcus aureus and coagulase-negative staphylococci in a community and a tertiary care hospital. J ClinMicrobiol42:2777-9.
- Wielders, C.L., Brisse, S., Graaf-Miltenburg, L. A., Troaelstra, A., Fleer, A., Schmitz, F. J.(2001).Evidence for in-vivo transfer of mecA DNA between strains of Staphylococcusaureus. The Lancet. 357: 9296-9272.



