Structural Analysis and 3D-Modeling of FANCD2 Proteins
Abubaker H. Mohamed1, AbdAlla A. AbdAlla2, Abdalbasit Mariod3, 4, *
1Department of Bioinformatics, Africa City of Technology, Khartoum, Sudan
2Department of Biotechnology, Omdurman Islamic University, Omdurman, Sudan
3Department of Biology, Faculty of Sciences and Arts-Alkamil, Jeddah University, Alkamil, Saudi Arabia
4Food Science & Technology Department, College of Agricultural Studies, Sudan University of Science & Technology, Khartoum North, Sudan
Abstract
Fanconi anemia (FA), is an uncommon inherited disease, is connected with dynamic bone marrow failure, inclination to cancer, and genomic instability. FA described by formative deformities, short stature, bone marrow failure, and a high danger of cancer. The mRNA of FANCD2 was recovered from the NCBI database, then interpreted into 679 amino acid by GeneMark server and study capacity of Fanconi anemia ID complex (FANCI-FANCD2) in light of the SWISSPORT Blast report: assumes a focal part in the repair of DNA interstrand cross-links, activated by means of DNA damage-induced phosphorylation by ATR and monoubiquitination by the FA center complex ubiquitin ligase, this capacity that permit processing and evacuation of crosslinked DNA and subsequently advances cell survival after DNA damage. The preservation homology was assessed by different arrangement utilizing BioEdit pakage, demonstrating high characters with another FANCD2 protein H. sapiens. The study structural and functional characterization, molecular formula, pI, EC, AI, GRAVY, instability index, half-life time and secondary structure were anticipated, this outcome affirm FANCD2 protein is steady, which can be utilized to confine and study in the vitro. In the present study a complete structural investigation and 3-D demonstrating of the FANCD2 protein utilized CPHmodels 3.2 server by utilizing template pdb of Mus musculus model because of the high rate of character (71.7%) and envision by Chimera 1.8. programming. Anticipated model was further surveyed by VERIFY-3D, this confirmation of quality of model.
Keywords
Fanconi Anemia, Protein, 3-D Modeling, GeneMark Server, SWISSPORT Blast Report, Structural and Functional Characterization
Received: August 8, 2015
Accepted: August 22, 2015
Published online: September 7, 2015
@ 2015 The Authors. Published by American Institute of Science. This Open Access article is under the CC BY-NC license. http://creativecommons.org/licenses/by-nc/4.0/
1. Introduction
Fanconi anemia (FA) is a complex infection that is viewed as a congenital type of aplastic anemia, the genetic method of transmission is both autosomal and X-linked, and a developing number of recognized genes are disseminated among the different chromosomes. The most widely recognized clinical appearance in many patients with FA, which may happen in all FA patients eventually, is a life-threatening bone marrow failure (BMF).1-2 The FA is also associated with diverse birth defects and a predisposition to malignancies. FA-related congenital malformations can influence numerous organ systems, including the central nervous system, the gastrointestinal system, and the skeletal system. 3–8 Other outcomes in patients with FA incorporate short stature, skin pigmentation variations from the norm, and small facial features. Furthermore, more than 70% of patients with FA show endocrine dysfunctions incorporating lacks in development hormone and thyroid hormone and in addition diabetes. 9-10 All of these disease manifestations recommend a part for FA genes in mechanisms that bear on hematopoiesis, development, and neoplasia. Fanconi anemia (FA) is an uncommon genome-instability, disorder with the variable presence of cellular hypersensitivity to DNA-inter strand cross connecting agents 11. There are no less than 12 of the basic genes have been recognized: FANCA, FANCB, FANCC, FANCD1/BRCA2, FANCD2, FANCE, FANCF, FANCG/XRCC9, FANCJ/BRIP1, FANCL/PHF9, FANCM/HEF, and FANCN/PALB2 12-16. Eight of the FA proteins (FANCA, -B, -C, -E, -F, -G, -L, and -M) and other parts collect in a nuclear complex, the FA "core complex," that is needed for the monoubiquitination of FANCD2 at amino acid residue K561 17-18. Monoubiquitination happens in response to DNA damage and amid the S phase of the cell cycle 18-19. The monoubiquitinated FANCD2 isoform (FANCD2-L, as opposed to FANCD2-S) is focused to nuclear foci containing proteins such as BRCA1, BRCA2, and RAD51 that are included in DNA-damage signaling and recombination repair 20-23. The FANCD2-lacking DT40 cells show imperfections in homologous recombination-mediated DNA double strand break (DSB) repair, translation synthesis, and gene conversion 20, 24, 25. Hence, FANCD2 is thought to assume a focal part in the support of genome stability 20, 25-26.
2. Materials and Methods
The mRNA of FANCD2 gene was retrieved from the NCBI database 27 GenBank: [AK022613] and translated into the protein by GeneMark software. The selected sequence for physiochemical analysis and 3-D modeling is given below:
NSDEINIPRLIVSQLKWLDRVVDGKDLTTKIMQLISIAPENLQHDIITSLPEILGDSQHA
DVGKELSDLLIENTSLTVPILDVLSSLRLDPDFLLKVRQLVMDKLSSIRLEDLPVIIKFI
LHSVTAMDTLEVISELREKLDLQHCVLPSRLQASQVKLKSKGRASSSGNQESSGQSCIIL
LFDVIKSAIRYEKTISEAWIKAIENTASVSEHKVFDLVMLFIIYSTNTQTKKYIDRVLRN
KIRSGCIQEQLLQSTFSVHYLVLKDMCSSILSLAQSLLHSLDQSIISFGSLLYKYAFKFF
DTYCQQEVVGALVTHICSGNEAEVDTALDVLLELVVLNPSAMMMNAVFVKGILDYLDNIS
PQQIRKLFYVLSTLAFSKQNEASSHIQDDMHSVIRKQLSSTVFKYKLIGIIGAVTMAGIM
AADGSESPSLTQERANLSDEQCTQVTSLLQLVHSCSEQSPQASALYYDEFANLIQHEKLD
PKALEWVGHTICNDFQDAFVVDSCVVPEGDFPFPVKALYGLEEYDTQDGIAINLLPLLFS
QDFAKDGGPVTSQESGQKLVSPLCLAPYFRLLRLCVERQHNGNLEEIDGLLDCPIFLTDL
EPGEKLESMSAKERSFMCSLIFLTLNWFREIVNAFCQETSPEMKGKVLTRLKHIVELQII
LEKYLAVTPDYVPPLGNFD
The capacity of the FANCD2 protein through conserved domain (CD) for indistinguishable or exceptionally comparable from BLASTp (protein-protein BLAST) Algorithm and utilized (SWISSPROT) database 28-29. The Multiple Sequence Alignment (MSA) by BioEdit package 30. Total 3 sequences that have higher comparability with a low E - value of BLASTp output, was taken for this study.
Physiochemical characterization, theoretical isoelctric point (pI), aggregate number of positive and negative deposits, extinction coefficient, instability index, half life time, aliphatic index and grand average hydopathy (GRAVY) were computed utilizing Expasy’s Prot-Param server 31, this apparatuses which permits the computation of different physical and chemical parameters for a given protein stored in Swiss-Prot or TrEMBL or for a user entered sequence.
The secondary basic components of FANCD2 protein was considered for this study that was utilized in SOPMA view 32, another exceptionally exact optional structure prediction method, SOPMA Expasy’s incorporates two feed-forward neural networks, which perform an investigation on yield got from every one of the successions of an arrangement of adjusted proteins fitting in with the same group of the database.
The 3-dimensional structure expectation we utilized CPHmodels 3.2 server to predict the 3D structure of those proteins with an unknown 3D structure model 33, 34. It is a protein homology modeling server, where the template recognition is based on profile-to-profile arrangement, guided by secondary structure and presentation forecasts. Visualization and characterization of the FANCD2 protein model were done by CHIMERA software and structural validation of protein model was done 35. Further, the model was surveyed by VARIFY 3D 36 which determine stereochemical viewpoints alongside principle chain and side chain parameters with a complete examination.
3. Results and Discussion
In this study the FANCD2 protein sequence of H. sapiens was translated from mRNA of FANCD2 gene taken from NCBI database, after then the protein sequence was checked for their function with a conserved domain (CD) utilizing pBLAST analysis (Table 1). This protein has domain function: CD [11721], Fanconi anemia D2 protein; The Fanconi anemia ID complex comprises of two subunits, Fanconi anemia I and Fanconi anemia D2 (FANCI-FANCD2) and plays a focal part in the repair of DNA interstrand cross-links (ICLs). The complex was activated via DNA damage-induced phosphorylation by ATR (ataxia telangiectasia and Rad3-related) and monoubiquitination by the FA core complex ubiquitin ligase, and it was bound to DNA at the ICL site, perceiving branched DNA structures. Defects in the complex causes Fanconi anemia, a cancer predisposition syndrome. The phosphorylation of FANCD2 was required for DNA damage-induced intra-S phase checkpoint and for cellular resistance to DNA cross linking agents.
Table 1. pBlast report of FANCD2 protein.
| Organism Name | Accession | % Identity | E value | Total score |
| Homo sapiens | Q9BXW9 | 99% | 0 | 1377 |
| Rattus norvegicus | Q6IV68 | 78% | 0 | 1074 |
| Mus musculus | Q80V62 | 77% | 0 | 1073 |
Multiple sequence alignment investigation of the FANCD2 protein was completed by BioEdit package that cover 3 sequences of pBLAST yield document in light of the fact that these sequences showed interesting similitude and low e-value. Predicted MSA had demonstrated high ration area (black and gray color) and low variable regions took after by color coding as red, blue and green (Fig. 1). The outcome was indicating over 99%, 78% and 77% identities with Homo sapiens, Rattus norvegicus and Mus musculus, respectively and can support the conservation function of FANCD2 protein.

Fig. 1. Multiple Sequence Alignment of FANCD2 protein.
Fig. 2. Secondary structure of the FANCD2 protein.
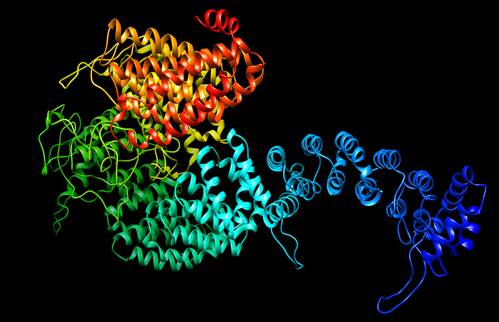
Fig. 3. 3-D model of FANCD2 protein.
Table 2. Total amino acid in FANCD2 protein.
| Amino acid | Number | Composition (%) |
| Ala (A) | 35 | 5.2 |
| Arg (R) | 21 | 3.1 |
| Asn (N) | 22 | 3.2 |
| Asp (D) | 44 | 6.5 |
| Cys (C) | 15 | 2.2 |
| Gln (Q) | 38 | 5.6 |
| Glu (E) | 43 | 6.3 |
| Gly (G) | 27 | 4.0 |
| His (H) | 15 | 2.2 |
| Ile (I) | 52 | 7.7 |
| Lue (L) | 95 | 14.0 |
| Lys (K) | 38 | 5.6 |
| Met (M) | 14 | 2.1 |
| Phe (F) | 28 | 4.1 |
| Pro (P) | 25 | 3.7 |
| Ser (S) | 67 | 9.9 |
| Thr (T) | 30 | 4.4 |
| Trp (W) | 4 | 0.6 |
| Tyr (Y) | 17 | 2.5 |
| Val (V) | 49 | 7.2 |
| Pyl (O) | 0 | 0.0 |
| Sec (U) | 0 | 0.0 |
The predicted physiochemical characteristic of the FANCD2 protein demonstrated that the molecular formula of FANCD2protein was C3409H5452N874O1028S29, have 10792atom that provided the ab-into information of this protein. The amino acid composition of the FANCD2 protein has 87(Asp + Glu) negatively charged, 59 (Arg+ Lys) positively charged, these outcome significance acidic nature of FANCD2 protein (Table.2). The genuine predicted molecular weight of the FANCD2 protein was 76059.7 Da, which was valuable in the isolation of this protein with the western blot procedure. The computed pI value was 4.98 and was less than 7 (pI<7) demonstrated that target protein was considered as acidic. This additionally corresponds with the number of negatively charged residues presented in the FANCD2 protein. At isoelectric point (pI), proteins were stable and compact so the computed pI will be valuable for building up a buffer system for purification by isoelectric centering technique and in pH based separation. Expasy’s ProtParam computed the extinction coefficient at 280 nm measured in water was 48205 M-1 cm-1. The extinction coefficient demonstrated the presence of Cys residues were diminished, such as low concentrations of Trp, Met and Tyr, that help us in the quantitative investigation of the protein-protein and protein-ligand interactions in solution.
3.1. Half Lifetime, Stability and Solubility
The evaluated half life time of FANCD2 protein was figured to be 1.4 hours (mammalian reticulocytes, in vitro), 3 min (yeast, in vivo) and less than 10 hours (Escherichia coli, in vivo) by N-end rule 30, which related the half-life of a protein to the identity of its N-terminal residue; the prediction was given for 3 model organisms (human, yeast and E. coli). The anticipated half life time demonstrated that FANCD2 was a stable protein in the cell after synthesis. The instability index of the FANCD2 protein was calculated (in vitro) to be 46.17. A protein whose instability index was smaller than 40 was predicted as stable, a worth over 40 predicts that the protein may be unstable 11. Therefore, the ascertained result has been demonstrated that FANCD2 protein was unstable in nature and this outcome was also correlated with calculated half life time value. The aliphatic index of the FANCD2 protein was predicted to be very high as 110.52, demonstrated that it may sustain on a wide temperature range because of its positive factor for the increment of thermal stability of globular proteins because of their relative volume involved in the aliphatic side chain (alanine, Valine, Isoleucine, and leucine). The computed GRAVY of FANCD2 protein was 0.074which demonstrated the hydrophilic nature of protein that supports the soluble nature of the FANCD2 protein, which can solve the frontier problem of isolation of soluble protein because acquiring of high-concentrations of soluble proteins remains an intensive experimental challenge 24.
3.2. Secondary and Tertiary Structure
The secondary structures of proteins are the frequently repeating local structures stabilized by hydrogen bonds. The most widely recognized samples were the alpha-helix and beta-sheet. It revealed that 447 were helices (65.83%), 18 were strands (2.65%), 12 were Beta (1.77%) and 202 were coils (29.75%) presented at different positions in the FANCD2 protein structure of H. sapiens (Fig.2). The outcome revealed that random helices were dominated among the secondary structure.
The three dimensional structure prediction of the FANCD2 protein (Fig.3) was built by utilizing template pdb of Mus musculus model (pdb id 3S4W) because of the high percentage of identity (71.7%), score (969) and e-value (0.0), because a high level of sequence identity should guarantee a more accurate alignment between the targeted sequence and template structure. The model was constructed for the target protein using CPH model 3.2 servers 37 to obtain PDB file and visualize by Chimera 1.8.
The quality of predicted structure of the four was further surveyed and affirmed by VARIFY 3D. The scores (from -1 to +1) were added and plotted for individual residues. The residues falling in the area where the orange line crosses 0.0 have low prediction accuracy and less stable conformation whereas, most of the residues fall above -0.17-0.71 so we can say that the model is of good quality (Fig. 4).
4. Conclusions
The FANCD2 protein is very important in the repair and damage DNA. The overall results provided the evidences that the FANCD2 protein of H. sapiens was a stable protein having acidic nature, it can be utilized to detect Fanconi anemia via Western Blot.
References
- G. C. Bagby, J. M. Lipton, E. M. Sloand, C. A. Schiffer, Marrow failure, Hematology, (Am Soc Hematol Educ Program). 318–336 (2004).
- M. Buchwald and M. Carreau "Genetic basis of Fanconi anemia," in Aplastic Anemia, H. Schrezenmeier and A. Bacigalupo, Eds., p. 403, Cambridge University Press, Cambridge UK, (1999).
- P. F. Giampietro, Verlander P. C., J. G. Davis, A. D. Auerbach "Diagnosis of Fanconi anemia in patients without congenital malformations: an international Fanconi anemia Registry Study," American Journal of Medical Genetics. 68 (1):58–61(1997).
- P. F. Giampietro, B. Adler-Brecher, P. C. Verlander, S. G. Pavlakis, J. G. Davis A. D. Auerbach. The need formore accurate and timely diagnosis in Fanconi anemia: a report from the international Fanconi anemia registry," Pediatrics, 91(6):1116–1120 (1993).
- E. T. Tsilou, N. Giri, S. Weinstein, C. Mueller, S. A. Savage, B. P. Alter, Ocular and orbital manifestations of the inherited bone marrow failure syndromes: Fanconi anemia and dyskeratosis congenita," Ophthalmology, 117, (3):615–622 (2010).
- M. J. Vale, M. J. Dinis, M. Bini-Antunes, B. Porto, J. Barbot, M. B. Coutinho Audiologic abnormalities of Fanconi anaemia, Acta Oto-laryngologica, 128( 9): 992–996 (2008).
- M. Tischkowitz and I. Dokal, Fanconi anaemia and leukaemia—clinical and molecular aspects, British Journal of Haematology, 126 (2):176–191(2004).
- N. Sari, C. Akyuz, D. Aktas, Wilms tumor, AML and medulloblastoma in a child with cancer prone syndrome of total premature chromatid separation and Fanconi anemia," Pediatric Blood and Cancer, 53(2): 208–210(2009).
- M. P Wajnrajch, J. M. Gertner, Z. Huma, Evaluation of growth and hormonal status in patients referred to the international Fanconi anemia registry, Pediatrics, 107(4 I): 744–754 (2001).
- N. Giri, D. L. Batista, B. P. Alter, C. A. Stratakis, Endocrine abnormalities in patients with Fanconi anemia, Journal of Clinical Endocrinology and Metabolism, 92( 7): 2624–2631(2007).
- H. Joenje, K.J. Patel, The emerging genetic and molecular basis of Fanconi anaemia. Nature Reviews Genetics. 2: 446–457(2001).
- S. Reid, D. Schindler, H. Hanenberg, K. Barker, S. Hanks, R. Kalb, K. Neveling, P. Kelly, S. Seal, M. Freund, Biallelic mutations in PALB2 cause Fanconi anemia subtype FA-N and predispose to childhood cancer. Nature Genetics, 39:162–164 (2007).
- B. Xia, J.C. Dorsman, N. Ameziane, Y. de Vries, M.A. Rooimans, Q. Sheng, G. Pals, A. Errami, E. Gluckman, J. Llera, Fanconi anemia is associated with a defect in the BRCA2 partner PALB2. Nature Genetics, 39:159–161(2007).
- O. Levran, C. Attwooll, R.T. Henry, K.L. Milton, K. Neveling, P. Rio, S.D. Batish, R. Kalb, E. Velleuer, S. Barral, The BRCA1-interacting helicase BRIP1 is deficient in Fanconi anemia. Nature Genetics, 37:931–933(2005).
- A.R. Meetei, A.L. Medhurst, C. Ling, Y. Xue, T.R. Singh, P. Bier, J. Steltenpool, S. Stone, I. Dokal, C.G. Mathew, A human ortholog of archaeal DNA repair protein Hef is defective in Fanconi anemia complementation group M. Nature Genetics, 37:958–963(2005).
- M. Levitus, Q. Waisfisz, B.C. Godthelp, Y. de Vries, S. Hussain, W.W. Wiegant, E. Elghalbzouri-Maghrani, J. Steltenpool, M.A. Rooimans, G. Pals, The DNA helicase BRIP1 is defective in Fanconi anemia complementation group J. Nature Genetics, 37: 934–935(2005).
- A.R. Meetei, S. Sechi, M. Wallisch, D. Yang, M.K. Young, H. Joenje, M.E. Hoatlin, W. Wang, A multiprotein nuclear complex connects Fanconi anemia and Bloom syndrome. Molecular Cell Biology, 23: 3417–3426 (2003).
- I. Garcia-Higuera, T. Taniguchi, S. Ganesan, M.S. Meyn, C. Timmers, J. Hejna, M. Grompe, A.D. D’Andrea, Interaction of the Fanconi anemia proteins and BRCA1 in a common pathway, Molecular Cell Biology, 7: 249–262(2001).
- T. Taniguchi, I. Garcia-Higuera, P.R. Andreassen, R.C. Gregory, M. Grompe, A.D. D’Andrea, S-phase-specific interaction of the Fanconi anemia protein, FANCD2, with BRCA1 and RAD51. Blood, 100:2414–2420 (2002).
- L.H. Thompson, J.M. Hinz, N.A. Yamada, N.J. Jones, How Fanconi anemia proteins promote the four Rs: replication, recombination, repair, and recovery.Environmental and Molecular Mutagenesis, 45:128–142(2005).
- K. Nakanishi, Y.G. Yang, A.J. Pierce, T. Taniguchi, M. Digweed, A.D. D’Andrea, Z.Q. Wang, M. Jasin, Human Fanconi anemia monoubiquitination pathway promotes homologous DNA repair.PNAS, Proceedings of the National Academy of Sciences, 102:1110–1115(2005).
- W. Niedzwiedz, G. Mosedale, M. Johnson, C. Y. Ong, P. Pace, K. J. Patel, The Fanconi anaemia gene FANCC promotes homologous recombination and error-prone DNA repair. Molecular Cell, 15:607–620(2004).
- M. Digweed, S. Rothe, I. Demuth, R. Scholz, D. Schindler, M. Stumm, M. Grompe A. Jordan, K. Sperling, Attenuation of the formation of DNA-repair foci containing RAD51 in Fanconi anaemia. Carcinogenesis, 23:1121–1126(2002).
- K. Yamamoto, S. Hirano, M. Ishiai, K. Morishima, H. Kitao, K. Namikoshi, M. Kimura, N. Arakawa, J. M. Buerstedde, K. Komatsu, L. H. Thompson, M. Takata, Fanconi anemia protein FANCD2 promotes immunoglobulin gene conversion and DNA repair through a mechanism related to homologous recombination. Molecular Cell Biology, 25:34–43(2005).
- K.D. Mirchandani, A.D. D’Andrea. The Fanconi anemia/ BRCA pathway: a coordinator of cross-link repair. Experimental Cell Research. 312:2647–2653(2006).
- M. Levitus, H. Joenje, J.P. deWinter. The Fanconi anemia pathway of genomic maintenance, Cell Oncology, 28:3–29 (2006).
- http://www.ncbi.nlm.nih.gov retrieved on 12.07.2014
- S. F. Altschul, T. L. Madden, A. A. Schäffer, J. Zhang, Z. Zhang, W. Miller, D. J. Lipman "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Research, 25:3389-3402 (1997).
- S. F. Altschul, J. C. Wootton, E. M. Gertz, R. Agarwala, A. Morgulis, A. A. Schäffer, Y-K. Yu "Protein database searches using compositionally adjusted substitution matrices", Federation of European Biochemical Societies (FEBS), 272:5101-5109, (2005).
- T. Hall, BioEdit: a user- friendly biological sequence alignment editor and analysis program for windows 95/98/NT, Nucleic Acids Symposium. 41:95-98, (1999).
- E. Gasteiger, C. Hoogland, A. Gattiker, S. Duvaud, M.R. Wilkins, R.D. Appel, A. Bairoch, Protein Identification and Analysis Tools on the ExPASy Server. The proteomics Protocols Handbook, Human Press, 571-607, (2005).
- C. Geourjon, and G. Deléage, SOPMA: Significant improvement in protein secondary structure prediction by consensus prediction from multiple alignments.Computer Applications in the Biosciences (Cabios.) 11, 681-684. (1995).
- M. Nielsen, C. Lundegaard, O. Lund, T.N. Petersen, CPHmodels-3.0 - Remote homology modeling using structure guided sequence profiles, Nucleic Acids Research, 38, doi:10.1093/nar/gkq535. (2010).
- O. Lund, M. Nielsen, C. Lundegaard, P. Worning. CPHmodels 2.0: X3M a Computer Program to Extract 3D Models, Abstract at the CASP5 conferenceA102, (2002).
- E. F. Pettersen, T.D. Goddard, C.C. Huang, G.S. Couch, D.M. Greenblatt, E.C Meng, T.E. Ferrin, UCSF Chimera--a visualization system for exploratory research and analysis. Journal of Computing Chemistry, 25(13):1605-12. (2004).
- D. Eisenberg, R. Luthy, J. U. Bowie, VERIFY3D: assessment of protein models with three dimensional profiles, Methods in Enzymology, 277: 396‐404 (1997).
- http://www.cbs.dtu.dk/services/CPHmodels retrieved on 15.07.2014.



