Phylogenetic Relationship Among Apicomlexan Parasites Based on In silico Analysis of Enzymes of the MEP Pathway
Shiv Kumar1, Ashish Kumar Gupta2, Deepak Ganjewala2, *
1Central Drugs Standard Control Organization (East Zone), Ministry of Health and Family Welfare, Government of India, Nizam Palace, Kolkata, India
2Amity Institute of Biotechnology, Amity University, Sector 125, Uttar Pradesh, India
Abstract
Apicomplexans such as P. falciparum use MEP pathway to synthesize isoprenoids crucial for their survival in the host. Here we report phylogenetic relationship among 11 parasites of phylum apicomplexa based on in silico analysis of enzymes of the MEP pathway with reference to Plasmodium falciparum 3D7. In addition, structure based homology of DXR of apicomlexans were performed with respect to DXR of Mycobacterium tuberculosis in order to gain detail insight of their DXR structure. The study revealed that 9 of 11 apicomlexans showed presence of the MEP pathway and all of its enzymes while the MEP pathway was absent in two members namely Cryptosporidium hominis and C. parvum. In two apicomlexans Toxoplasma gondi and Eimeria tenella an enzyme MCT and in P. knowlesi HDR enzyme of the MEP pathway was absent. Sequence analysis of the MEP pathway enzymes and resultant cladogram indicated that plasmodium sp. can be easily distinguished from non-plasmodium sp. The cladogram revealed close relationship among plasmodium sp. viz., P. falciparum 3D7, P. berghei str. ANKA, P. vivax SaI-1 and P. knowlesi H. Results of structure based homology analysis showed presence of amino acids viz., Thr (T) 21, Gly (G) 22, Ser (S) 23, Ile (I) 24, Gly (G) 47, Gly (G) 48, Ala (A) 49 and Glu (E) 129 in the NADPH binding domain; Asp (D)151, Glu (E)153 and Glu in Mn2+ binding domain and Ser (S)152, Ser (S) 177, His (H) 200, Asn (N) 218 and Lys (K)219 in fosmidomycin binding domain in DXRs of apicomlexans. These amino acids were found to be highly conserved. Thus, MEP pathway enzymes served as excellent tools to discern phylogenetic relationship in apicomlexans and an attractive target for development of new anti-parasitic drugs against these parasitic microorganisms.
Keywords
Apicomplexa, Apicoplast, Fosmidomycin, Isoprenoid, MEP Pathway, Plasmodium falciparum
Received: June 30, 2015
Accepted: July 20, 2015
Published online: August 2, 2015
@ 2015 The Authors. Published by American Institute of Science. This Open Access article is under the CC BY-NC license. http://creativecommons.org/licenses/by-nc/4.0/
1. Introduction
The Apicomplexa group of protozoa comprises approximately 5000 species of which majority causes infectious diseases in animals (Cavalier-Smith 1993). Important species of this group such as Plasmodium falciparum, P. vivax, P. berghei and P. knowlesi are well known as they are causative agents of malaria in human. Among others, Toxoplasma gondii causes toxoplasmosis; Babesia bovis Texas Cattle Fever and Theileria parva is the causative agent of East Coast fever, corridor disease, and Zimbabwean theileriosis (Wasmuth et al., 2009). Table 1 summarizes host and environment specificity for apicomplexans used in this study. Members of the apicomplexa are characterized by the presence of four-membrane relict plastid called apicoplast (Arora et al., 2010). Apicoplast is a 35 KB extra chromosomal DNA in P. falciparum which harbor around 551 nucleus encoded apicoplast targeted (NEAT) proteins (Gardner 2002). Many of these NEAT proteins are promising drug targets for killing of the Apicomplexan parasites (Steinbacher et al., 2002). Apicoplast is essential for survival of the apicomlexans as it possess many pathways of formation of fatty acids, heme and isoprenoids (Roos et al., 2002). Isoprenoids play crucial roles in the survival of malarial parasite P. falciparum. Isoprenoids are one of the major classes of secondary metabolites which consist of C5 compound called isoprenes or isopentenyl diphosphate (IPP) and its isomer dimethyl allyl diphosphate (DMAPP). In plants and many bacteria isoprenoids are biosynthesized via 2C-methyl-D-erythritol-4-phosphate (MEP) pathway (Rohmer et al., 1993; Eisenreich et al., 1998; Ganjewala et al., 2009). Apicomlexans also have the MEP pathway in apicoplasts. However, the enzymes of this pathway are being imported from the nucleus as NEAT proteins/enzymes (Kemp et al., 2002). The genes encoding enzymes of the MEP pathway have been isolated and characterized from Plasmodium species (Kemp et al., 2002; Rohdich et al., 2001). The MEP pathway comprises seven enzymatic steps (Fig. 1). The first step is the formation of 1-deoxy-D-xylulose-5-phosphate (DOXP) by condensation of pyruvate and glyceraldehyde (Takahashi et al., 1998) catalyzed by 1-deoxy-D-xylulose-5-phosphate synthase (DXS). The second step DXP leads to formation of 2-C-methyl-erythritol-4-phosphate (MEP) from DOXP by an NADPH–dependent enzyme 1-deoxy-D-xylulose-5-phosphate reductoisomerase (DXR; EC 1.1.1.267) (Argyrou and Blanchard 2004). These two initial steps are the rate limiting or committed steps of the MEP pathway hence enzymes DXS and DXR appears an attractive target for development of enzyme based novel anti-parasitic drugs. The DXR of P. falciparum and P. vinckei is reported to be inhibited by fosmidomycin thereby arresting the formation of isoprenoids (Henriksson et al., 2007).
Currently, complete genome sequences of several apicomplexan parasites are available which helped to understand pathogenicity of apicomplexans at molecular level (Wasmuth et al., 2009; Ajioka et al., 1998; Howe 2001; Li et al., 2003a; Cui et al., 2005). The present study was undertaken to discern phylogenetic relationship among 11 parasite members of apicomplexan family by in silico analysis of the enzymes of the MEP pathway using P. falciparum 3D7 as a reference. In addition, structure based homology analysis of DXRs of apicomlexans were performed with respect to DXR of M. tuberculosis to gain detail insight in to amino acid residues of DXRs which interacts and binds with NADPH, Mn2+ and fosmidomycin.
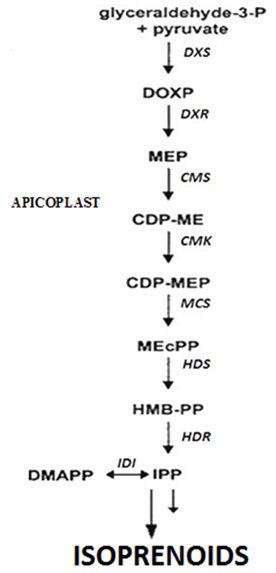
Figure 1. MEP pathway of isoprenoid biosynthesis. CDP-ME, 4-(cytidine-5'-diphospho)-2-C-methyl-D-erythritol; CDP-MEP, 4-diphosphocytidyl-2Cmethyl-D-erythritol 4-phosphate; DMAPP, dimethylallyl diphosphate; DOXP, 1-deoxy-D-xylulose 5-phosphate; GAP, glyceraldehyde 3-phosphate; GGPP, geranylgeranyl diphosphate; GPP, geranyl diphosphate; HBMPP, 4-hydroxy- 3-methylbut-2-enyl diphosphate; IPP, isopentenyl diphosphate; ME-cPP, 2Cmethyl-D-erythritol 2,4-cyclodiphosphate; MEP, 2C-methylerythritol 4-phosphate;Enzymes are: indicated in italic; CMK, 4-(cytidine-5'-diphospho)-2-C-methyl-D-erythritol kinase; CMS, 4-diphosphocytidyl-2Cmethyl- D-erythritol 4-phosphate synthase; DXR, 1-deoxy-D-xylulose 5-phosphate reductoisomerase; DXS, 1-deoxy-D-xylulose 5-phosphate synthase; HDS, 4-hydroxy-3-methylbut2-en-yl-diphosphate synthase; MCS, 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase; IDI, isopentenyl-diphosphate isomerise.
2. Materials and Methods
2.1. Retrieval of Enzymes Sequences
Database searches were performed independently for seven enzymes of the MEP pathway in selected Apicomlexans (Table 1). Completely annotated genomes are available online P. falciparum at PlasmoDB (release version 4.4; http://plasmodb.org/), T. gondii at ToxoDB (release version 3.0; http://www.toxodb.org/),C. parvum (version 3.4), and C. hominis (version 3.4) at ApiDB, T. parva and T. annulata at GenBank, NCBI (Wasmuth et al., 2009). BLAST algorithm was used for similarity searches in the above databases and sequence homology analysis was performed among seven enzymes of the MEP pathway retrieved from selected apicomlexans using specialized BLAST (bl2seq) from http://www.ncbi.nlm.nih.gov/blast/Blast.cgi server. Amino acid sequences of enzymes of the MEP pathway were obtained from NCBI, a publicly available database.
Table 1. Host and environment specificity of apicomplexa parasites selected for the study.
| Species | Definitive host | Intermediate host | Preferred host environment | Disease |
| Plasmodium falciparum | Mosquito | Human | Erythrocyte | Malaria |
| Plasmodium vivax | Mosquito | Human | Erythrocyte | recurring (tertian) malaria |
| Plasmodium berghei | Mosquito | Rat | Erythrocyte | Malaria |
| Plasmodium knowlesi strain H | macaques (monkeys) | humans | Erythrocyte | 70% Malaria in South East Asia |
| Plasmodium yoelii | Mosquito | Rat | Erythrocyte | Malaria |
| Babesia bovis | Tick | cattle | Erythrocyte | Hemolytic anemia known as Babesiosis |
| Theileria annulata | Tick | Bovine | Leukocyte | tropical theileriosis (Mediterranean theileriosis) |
| Theileria parva | Tick | Bovine | Leukocyte | East Coast fever ( theileriosis) in cattle |
| Eimeria tenella | Poultry | None | Intestinal tract | Hemorrhagic cecal coccidiosis in young poultry |
| Toxoplasma gondii | Feline | Warm-blooded animals | Broad range | Toxoplasmosis |
| Cryptosporidium hominis | Human | None | Intestinal track | Gastroenteritis and diarrhea characteristic of cryptosporidosis |
| Cryptosporidium parvum | Mammal | None | Intestinal track | Gastroenteritis and diarrhea characteristic of cryptosporidosis |
Table 2. Distribution of genes encoding enzymes of the MEP pathway in selected Apicomplexan.
| Apicomlexans | DXS | DXR | MCT | CMK | MDS | HDS | HDR |
| P. falciparum 3D7 | + | + | + | + | + | + | + |
| Pl. vivax SaI-1 | + | + | + | + | + | + | + |
| P. berghei ANKA | + | + | + | + | + | + | + |
| P. knowlesi strain H | + | + | + | + | + | + | - |
| T. gondii ME49 | + | + | - | + | + | + | + |
| T. annulata strain Ankara | + | + | + | + | + | + | + |
| T. parva strain Muguga | + | + | + | + | + | + | + |
| B. bovis T2Bo | + | + | + | + | + | + | + |
| E. tenella | + | + | - | + | + | + | + |
| C. parvum Iowa II | None of the enzymes of MEP pathway was found | ||||||
| C. hominis | None of the enzymes of MEP pathway was found | ||||||
(+) presence and (-) absence of a gene in the genome
2.2. In silico Analysis
Multiple sequence alignment was performed with the Clustal W program at EMBL (http://www.ebi.ac.uk/) and GeneDoc. Homology (identity percentage) in amino acid sequences of enzymes of the MEP pathway from selected Apicomplexans parasites were compared with that of P. falciparum 3D7 using BLAST algorithms at NCBI (http://www.ncbi.nlm.nih.gov/BLAST/). We randomly used an expectation (E-value) cut off of 10-3. The non-homologous entries were then subjected to BLASTP against the DEG for the identification of homologous sequences to essential genes at a cut off score of 10-10.
3. Results and Discussion
3.1. Distribution of the MEP Pathway Enzymes
Plasmodium falciparum 3D7 use MEP pathway located in apicoplast to form isoprenoids. The genes encoding enzymes of the MEP pathways have been cloned and characterized from P. falciparum 3D7. Apart from P. falciparum the MEP pathway is present in other members of the apicomlexa which suggests that these members also dependent on isoprenoids for their survival. We performed In silico analyses of the seven enzymes of the MEP pathway in 11 members of Apicomplexa with respect to P. falciparum 3D7 as a reference. The study revealed the presence of the MEP pathway and its enzymes in 9 of the 11 selected apicomplexans (Table 2). In C. parvum Iowa II and C. hominis the MEP pathway was absent thereby suggesting no roles of isoprenoids in their survival and/ or pathogenicity. In E. tenella an enzyme MCT was absent which may be due to its very low expression beyond the limit of detection. It is reported that the Isp encoded enzyme are expressed at relatively low levels in P. falciparum, which may affect their representation within the partial genome data sets. This pathway is important in the modification of transfer RNAs, suppressing premature stop codons and frame shifts in coding regions (Petrulloet al., 1983).
3.2. Sequence Analysis and Phylogenetic Relationship
Amino acid sequences of enzymes of the MEP were analysed using BLAST and compared to homologous enzymes of P. falciparum 3D7. Three BLAST bit score cut offs were used for identity percentage analysis. The results revealed that only two enzymes of the MEP pathway DXR and HDR from apicomlexans displayed significantly higher 50-80% and 50-75% amino acid sequence similarity, respectively with homologous enzymes of P. falciparum 3D7 (Fig. 2). Four of the MEP pathway enzymes namely DXR, CMS, MCS and HDR of P. berghi showed significant (70%) amino acid sequence similarity whereas DXS, CMK and HDS only 55-70% with homologous enzymes of P. falciparum 3D7. The MEP pathway enzymes DXS, DXR, CMS and HDR from T. annulata demonstrated comparatively less 50-70% amino acid sequence similarity with those of P. falciparum 3D7 enzymes. In B. bovis, only two enzymes DXR and HDR showed 58% and 50% amino acid sequence similarity with enzymes of the reference organisms. The DXRs of P. vivax and T. knowlesi had 75% sequence similarity with the reference organisms and MCS and HDS 60-65% while the CMK shared only 50% amino acid sequence similarity with the corresponding enzymes of P. falciparum 3D7. The MEP pathway enzymes in T. parva and T. gondi displayed low amino acid sequence similarity with reference organisms P. falciparum 3D7. Multiple sequence alignment analysis of enzymes of the MEP pathway performed with the ClustalW revealed close phylogenetic relationship within the plasmodium sp. of phylum apicomplexan and they could be easily distinguished from distantly related non plasmodium sp. (Fig. 3)
3.3. Structure Based Homology Analysis
Structure based homology analysis of DXR enzyme of the MEP pathway performed using DXR of M. tuberculosis Hominis revealed the amino acid residues present in the NADPH and Mn2+ binding domains as well as those interacts with fosmidomycin. The essentiality of DXR in M. tuberculosis Hominis has been demonstrated recently (Goble et al., 2007). M. tuberculosis DXR provides the most complete picture of interactions in the active site of enzyme. The DXR is a NADPH dependent enzyme catalyse rearrangement and reduction of 1-deoxy-D-xylulose 5-phosphate (DXP) to form 2-C-methyl-D-erythritol 4-phosphate (MEP). The reaction requires the presence of a divalent cations such as Mg2+, Co2+or Mn2+ (Takahashi et al.,, 1998). Fosmidomycin is a potent inhibitor of DXR and hence it can block the MEP pathway (Henriksson et al., 2007). It is reported to be active against the protozoan parasite P. falciparum in humans (Missinou et al., 2002; Lell et al., 2003) and P. vinckei in mice (Jomma et al., 1999).
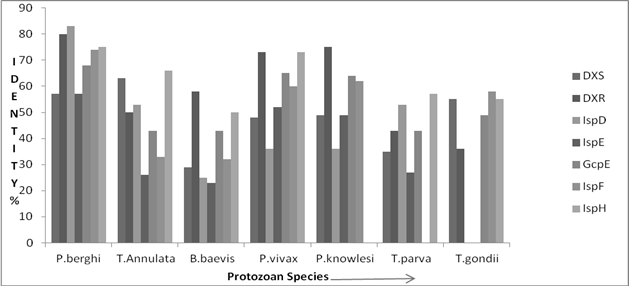
Figure 2. Reletive amino acid sequence similarity in enzymes of the MEP pathway with referene to P. falciparum 3D7.
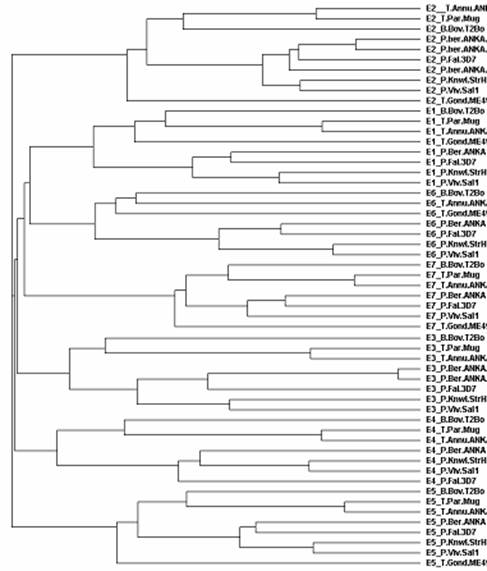

Figure 3. Cladogram depicting phylogenetic relationship among selected Apicomplexans. E1:DXS; E2: DXR; E3: CMS; E4:CMK; E5: HDS; E6: MCS; and E7: HDR.
Table 3. Conserved amino acid residues of NADPH and Mn+2 binding domains in crystal structure of DXR of M. tuberculosis and DXRs of Apicomplexans.
| Organisms/Apicomplexan | NADPH Binding Residues | Mn+2 Binding Residues | |||||||||
| M. tuberculosis | T21 | G22 | S23 | I24 | G47 | G48 | A49 | E129 | D151 | E153 | E222 |
| P. falciparum 3D7 | T | G | S | I | N | K | S | E | D | D | D |
| P. vivax SaI-1 | T | G | S | I | N | K | S | E | D | E | E |
| P. berghei str. ANKA | T | G | S | I | N | - | K | E | D | E | E |
| P. knowlesi strain H | T | G | S | I | N | K | S | E | D | E | E |
| T. gondii ME49 | T | G | S | I | G | G | S | E | D | E | E |
| T. annulata strain Ankara | T | G | S | I | K | S | N | E | D | E | E |
| T. parva strain Muguga | T | G | S | I | N | S | N | E | D | E | E |
| B. bovis T2Bo | T | G | S | I | N | D | E | E | |||
| E. tenella | T | G | S | I | G | D | E | E | |||
Earlier reports have revealed that in M. tuberculosis DXR, NADPH is bound in the N-terminal domain and comes in close contact with fosmidomycin and the Mn2+ ions that are positioned underneath the active site flap. Amino acid residues Thr (T) 21, Gly (G) 22, Ser (S) 23, Ile (I) 24, Gly (G) 47, Gly (G) 48, Ala (A) 49 and Glu (E) 129 of the N-terminal domain of M. tuberculosis DXR interact with NADPH. Amino acid residues Gly (G) 206 and Asn (N) 209 from the active site flap in the catalytic domain also interact with NADPH (Levitt and Perutz 1988). Structure based homology analysis performed here also revealed the presence of similar amino acid residues in the DXRs of apicomplexans which interacts with NADPH (Table 3A). Further the amino acid residues, Asp (D)151, Glu (E)153 and Glu (E)222 of DXR in M. tuberculosis were also present in the DXRs of apicomplexans. These amino acid residues interact with Mn2+ ion and fosmidomycin (Table 3). Fosmidomycin also interacts with amino acid residues, Ser (S)152, Ser (S) 177, His (H) 200, Asn (N) 218 and Lys (K)219. These amino acid residues found in the catalytic domain DXR of M. tuberculosis also elucidated in the DXRs of all apicomplexans studied (Table 4). Amino acids interacting with NADPH, Mn+2 and fosmidomycin deciphered from the crystal structure of DXR of M. tuberculosis Hominis are depicted in Fig. 4.
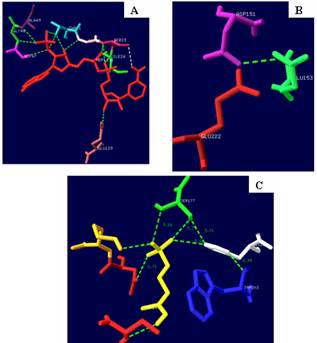
Figure 4. Amino acids interacting with [A] NADPH, [B] Mn+2 and [C] fosmidomycin deciphered from the crystal structure of DXR of M. tuberculosis Hominis.
Table 4. Conserved amino acid residues in crystal structure of DXR of M. tuberculosis and DXRs of Apicomplexans interacting with fosmidomycin.
| Organisms | Fosmidomycin Binding Pocket Residues in DXR | |||||
| 2JCV (M. Tuberculosis) | S152 | S177 | H200 | W203 | N218 | K219 |
| P. falciparum 3D7 | S | S | H | W | N | K |
| P. vivax SaI-1 | S | S | H | W | N | K |
| P. berghei ANKA | S | S | H | W | N | K |
| P. knowlesi H | S | S | H | W | N | K |
| T. gondii ME49 | S | S | H | W | N | K |
| T. annulata Ankara | S | S | H | W | N | K |
| T. parva Muguga | S | S | H | W | N | K |
| B. bovis T2Bo | S | S | H | W | N | K |
| E. tenella | S | S | H | W | N | K |
4. Conclusion
Apicomlexans parasites such as P. falciparum use MEP pathway to produce isoprenoids which are crucial for their survival and pathogenesis. Therefore the MEP pathway enzymes can be attractive targets for development of newer anti-parasite drugs. The MEP pathway enzymes may also serve as biochemical marker to elucidate phylogenetic relationship among species. In the present study, MEP pathway enzymes have been used as biochemical tools to discern phylogenetic relationship among the members of the apicomplexa family. A key enzyme DXR of the MEP pathway was subjected to structure based homology analyses with respect to DXR of M. tuberculosis. The results of these analyses have provided detail insight into amino acid composition of the NADPH, Mn2+ and fosmidomycin binding domains of the DXR. Most the amino acid residues were highly conserved. Although the outcome of the present study is preliminary but they may be useful for developing single anti-parasitic drug against broad spectrum of apicomlexan parasites rely on the MEP pathway for their survival.
Acknowledgments
Corresponding author of this article is grateful to Dr. Ashok Kumar Chauhan, Founder President and Mr. Atul Chauhan, Chancellor of Amity University, Uttar Pradesh, Noida, India for providing necessary support and facilities.
References
- Ajioka, J.W., Boothroyd, J.C., Brunk, B.P., Heh, A., Hillier, L., Manger, I.D. et al. (1998). Gene discovery by EST sequencing in Toxoplasma gondii reveals sequences restricted to the Apicomplexa. Genome Research, 8: 18-28
- Argyrou, A. and Blanchard, J.S. (2004). Kinetic and chemical mechanism of Mycobacterium tuberculosis 1-deoxy-D-xylulose-5-phosphate isomeroreductase. Biochemistry, 43:4375-4384
- Arora, N., Banerjee, A.K. and Murty, U.S.N. (2010). Homology model of 2-C-methyl-D-erythritol 2, 4-cyclodiphosphate (MECP) synthase of Plasmodium falciparum 3D7. Electronic Journal of Biology, 6: 52-57.
- Cavalier-Smith, T. (1993). Kingdom protozoa and its 18 phyla. Microbiology Review, 57: 953-994
- Cui, L., Fan, Q., Hu, Y., Karamycheva, S.A., Quackenbush, J., Khuntirat, B., Sattabongkot, J. and Carlton, J.M. (2005). Gene discovery in Plasmodium vivax through sequencing of ESTs from mixed blood stages. Molecular and Biochemical Parasitology, 144: 1-9
- Eisenreich, W., Schwarz, M., Cartayrade, A., Arigoni, D., Zenk, M.H. and Bacher, A. (1998). The deoxyxylulose phosphate pathway of terpenoid biosynthesis in plants and microorganisms. Chemical Biology, 5: R221-R233
- Ganjewala, D., Kumar, S. and Luthra, R. (2009). An account of cloned genes of methyl-erythritol-4-phosphate pathway of isoprenoid biosynthesis in plants. Current Issues in Molecular Biology, 11:S35-S45
- Gardener, M.J., Hall, N., Funq, E., White, O., Berriman, M., Hyman, R.W., et al (2002). Genome sequence of the human malaria parasite Plasmodium falciparum. Nature, 419:498-511
- Goble, J.L., Adendorff, M.R., de Beer, T.A.P., Stephens, L.L and Blatch, G.L. (2010). The malarial drug target Plasmodium falciparum 1-Deoxy-D-Xylulose-5-Phosphate reductoisomerase (PfDXR): Development of a 3-D model for identification of novel, structural and functional features and for inhibitor screening. Protein Peptide Letters, 17:109-120
- Henriksson, L.M., Unge, T., Carlsson, J., Aqvist, J., Mowbray, S.L. and Jones TA (2007) Structures of Mycobacterium tuberculosis 1-Deoxy-D-xylulose-5-phosphate reductoisomerase provide new insights into catalysis. Journal of Biological Chemistry, 282:19905-91916
- Howe, D.K. (2001). Initiation of a Sarcocystis neurona expressed sequence tag (EST) sequencing project: A preliminary report. Veterinary Parasitology, 95: 233-239
- Jomaa, H., Wiesner, J., Sanderbrand, S., Altincicek, B., Weidemeyer, C., Hintz, M., Turbachova, I., Ebert, M., Zeidler, J., Lichtenthaler, H.K., Soldatri, D. and Beck, E. (1999). Inhibitors of the nonmevalonate pathway of isoprenoid biosynthesis as antimalarial drugs. Science, 285: 1573-1576
- Kemp, L.E., Bond, C.S. and Hunter, W.N. (2002). Structure of 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase: an essential enzyme for isoprenoid biosynthesis and target for antimicrobial drug development. Proceeding of National Academy of Sciences USA, 99: 6591-6596
- Lell, B., Ruangweerayut, R., Wiesner, J., Missinou, M.A., Schindler, A., Baranek, T., Hintz, M., Hutchinson, D., Jomaa, H. and Kremsner, P.G. (2003). Fosmidomycin, a novel chemotherapeutic agent for malaria. Antimicrobial Agents Chemotherapy, 47: 735-738
- Levitt, M. and Perutz, M.F. (1988). Aromatic rings act as hydrogen bond acceptors. Journal of Molecular Biology, 201: 751-754
- Li, L., Brunk, B.P., Kissinger, J.C., Pape, D., Tang, K., Cole, R.H., et al., (2003). Gene discovery in the apicomplexa as revealed by EST sequencing and assembly of a comparative gene database. Genome Research, 13: 443-454
- Missinou, M.A., Borrmann, S., Schindler, A., Issifou, S., Adegnika, A.A., Matsiegui, P.B., et al., (2002). Fosmidomycin for malaria. Lancet, 360: 1941-1942
- Petrullo, L.A., Gallagher, P.J. and Elseviers, D. (1983).The role of 2-methylthio- N 6-isopentenyladenosine in readthrough and suppression of nonsense codons in Escherichia coli. Molecular and General Genetics, 190: 289-294
- Rohdich, F., Eisenreich, W., Wungsintaweekul, J., Hecht, S., Schuhr, C.A. and Bacher, A. (2001). Biosynthesis of terpenoids. 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (IspF) from Plasmodium falciparum. European Journal of Biochemistry, 268: 3190-3197
- Rohmer, M., Knani, M., Simonin, P., Sutter, B. and Sahm, H. (1993). Isoprenoid biosynthesis in bacteria: a novel pathway for the early steps leading to isopentenyl diphosphate. Biochemical Journal, 295: 517-52
- Roos, D.S., Crawford, M.J., Donald, R.G., Fraunholz, M., Harb, O.S., He, C.Y., Kissinger, J.C., Shaw, M.K. and Striepen, B. (2002). Mining the Plasmodium genome database to define organellar function: what does the apicoplast do?’, Philosophical Transaction of the Royal Society London B: Biological Science 357: 35-46
- Steinbacher, S., Kaiser, J., Wungsintaweekul, J., Hecht, S., Eisenreich, W., Gerhardt, S., Bacher, A. and Rohdich, F. (2002). Structure of 2- C -methyl-D-erythritol-2,4-cyclodiphosphate synthase involved in mevalonate independent biosynthesis of isoprenoids. Journal of Molecular Biology, 316: 79-88
- Takahashi, S., Kuzuyama, T., Watanabe, H. and Seto, H. (1998). A 1-deoxy-D-xyulose 5-phosphate reductoisomerase catalyzing the formation of 2-C-methyl-D-erythritol 4-phosphate in an alternative non mevalonate pathway for terpenoid biosynthesis. Proceedings of National Academy of Sciences USA, 95: 9879-9884
- Wasmuth, J., Daub, J., Peregrın-Alvarez, J.M., Finney, C.A.M. and Parkinson, J. (2009). The origins of apicomplexan sequence innovation. Genome Research, 19:1202-1213



