Avian Influenza Resistant Gene (Mx) and Its Diversity in Chicken and Duck
Minakshi Nag1, Md. Mostafizer Rahman1, Anjuman Ara Bhuyan2, Mridha Md. Kamal Hossain2, Md. Abdul Alim2, Md. Saidul Islam2, Mir Rowsan Akter1, Mahmudul Hasan3, Jahangir Alam2, *
1Hajee Mohammad Danesh Science and Technology University, Dinajpur, Bangladesh
2National Institute of Biotechnology, Ganakbari, Ashulia, Savar, Dhaka, Bangladesh
3Bangladesh Livestock Research Institute, Savar, Dhaka, Bangladesh
Abstract
Poultry sector has been facing threat of highly pathogenic avian influenza (HPAI) since 2007 in Bangladesh. The chicken Mx protein has been reported to exhibit antiviral activity against influenza virus. Therefore, targeting Mx gene may be an approach for development of AI resistant poultry. However, there is scarcity of information about the Mx gene and its diversity in chicken and duck in Bangladesh. To enumerate Mx gene and its diversity a total of 60 blood samples were collected from six chicken groups (Rhode Island Red, White Leghorn, White Rock, Barred Plymouth Rock, Necked Neck & Hilly) and four duck groups (Pekin, Rupali, Nageshwari & Common Deshi) from Bangladesh Livestock Research Institute, Nageshwary, Kurigram district and Natore Sadar, Natore district. Two sets of primers were used. Of them one set for Mx gene detection by PCR and another set for Mx gene detection and its diversity analysis by polymerase chain reaction restriction fragment length polymorphism (PCR-RFLP) using RsaI and SspI restriction enzymes. The Mx gene detection with one set primer was found to vary from 60.0-100% (average 77.8%) in chicken and 0-40% (average 43.6) in duck. However, with another set of primer detection rate was 40-100 and 66.7-100% in chicken and duck, respectively. PCR-RFLP analysis of 27 samples revealed that tested samples contain homozygous resistant (R/R), homozygous sensitive (S/S) and heterozygous (R/S) Mx allelic gene. In chicken the R/R, S/S and R/S Mx allelic gene was 38.9, 33.3 and 27.0% respectively; while in duck it was 11.1, 66.7 and 22.2%, respectively. Ducks were found more sensitive then tested chicken. Mx gene diversity was found to exist not only among the groups but also within the group of chicken and duck. To the best of our knowledge this is the first report on Mx gene study in Bangladesh covering both chicken and duck.
Keywords
Mx Gene, Diversity, Chicken, Duck, Avian Influenza
Received: April 9, 2015
Accepted: April 23, 2015
Published online: May 28, 2015
@ 2015 The Authors. Published by American Institute of Science. This Open Access article is under the CC BY-NC license. http://creativecommons.org/licenses/by-nc/4.0/
1. Introduction
Type I interferon (IFN) produced by virally-infected cells play a crucial role in the host’s early defence against viruses by helping in expression of a number of host genes, including Myxovirus resistant gene (Mx) (Muller et al., 1994; Horisberger, 1995). The specific function of the Mx protein in defence against viruses is the result of a direct interaction between the carboxyl terminus of the Mx protein of specific species of pathogenic viruses (Lee and Vidal 2002). Mx protein has been found in many organisms, including yeast, mouse, fish, birds, and mammals (Aebi et al., 1989; Meier et al., 1990; Rothman et al., 1990; Bazzigher et al., 1993). Most species have two Mx genes which code for proteins that accumulate in either the nucleus or the cytoplasm of cell. Mouse and rat Mx1 proteins are located in the nucleus, whereas most other Mx proteins are found in the cytoplasm (Haller et al., 2007). The nuclear mouse Mx1 protein primarily inhibits the replication of Orthomyxoviruses, including influenza viruses (Garber et al., 1991; Haller et al., 1995) and the cytoplasmic Mx2 protein mainly inhibits the vesicular stomatitis virus (Zurcheret al., 1992; Jin et al., 1999). Cytoplasmic Mx proteins such as the human MxA or bovine Mx1 not only confer antiviral activity against influenza viruses but also inhibit many unrelated viruses (Pavlovic et al., 1992; Schneider et al., 1994; Schwemmle et al., 1995; Landis et al., 1998; Baise et al., 2004). The Mx gene was first identified in fowl in 1980 when research on Mx genes revealed host defence mechanisms against influenza virus infections (Livant et al., 2007). In duck and chicken, only one Mxprotein was identified. The chicken Mx protein exhibit antiviral activity against the influenza virus and the recombinant vesicular stomatitis virus (VSV) (Ko et al., 2002). It is also reported that naturally, native chicken has ability to resist the virus controlled by antiviral Mx gene (Sartika et al., 2011). However, Benfield et al., (2008) reported that breed Shamo, which has an Asparagine at amino acid 631 is void of activity against the H5N1. Each chicken’s ability to resist the virus differs from one chicken to another and the antiviral gene control it. In chickens, the Mx gene is located in chromosome 1 in a 20767 bp fragment. It consists of 13 exons, with as many as 2115bp coding regions and 705 amino acids. Resistance against the AI virus was found at exon 13, nucleotide number 1892 where it undergoes alkaline transition mutation (single mutation) (Li et al., 2007). Mutations in the Mx gene cause triple codon changes to 631, which mutates the amino acid asparagine (AAC/AAU) to serine (AGC/AGU). Ko et al., (2002) showed that in many natural variations of chicken Mx gene, only the 631 (Ser to Asn) mutations was caused by a single nucleotide substitution in 2,032 point, which has antiviral activity. Chicken Mx gene is highly polymorphic, and that a single-nucleotide polymorphism affecting amino acid 631 determines antiviral activity (Janzen et al., 2000). Therefore a mutation of the amino acid asparagine to serine, the resulting polymorphism in the Mx gene will cause to be susceptible to AI virus subtype H5N1 and AI virus attacks (Li et al., 2007).
Livestock is an integral part of agricultural economy of Bangladesh. It provides food, nutrition, employment opportunity, income, draft power, fuel, manure, transport etc. Bangladesh Economic Review (2009) reported highest growth rate in livestock subsector than crop and fisheries. However, a declining growth rate was reported during 2006-2007 and 2007-2008 due to serious outbreak of highly pathogenic avian influenza (HPAI) in poultry. Native chicken (~140 million) and ducks (~38 million) are two important species of poultry distributed all over the country, source of protein among low income families, (Bhuiyan et al., 2005& 2013), predominantly reared by women thus enhancing women empowerment. With the emergence of HPAI H5N1 in Bangladesh in 2007, conventional control strategies based on surveillance, stamping out, movement restriction and enforcement of biosecurity measures did not prevent the virus spreading and outbreak. Moreover, a new clade 2.3.2.1 emerged in January 2011. Later government of Bangladesh decided to allow vaccination on experimental basis in 2012 and still going on. To aid in HPAI control program, alternative and complementary ways of controlling of HPAI can be explored. The use of antiviral chemotherapy and natural compounds, avian-cytokines, RNA interference, genetic breeding and/or development of transgenic poultry may be the options (Abdelwhab and Hafez 2012). Mx gene may be the target one for genetic breeding and/or development of AIV resistant bird. Thus determination of Mx gene and its diversity in chicken and duck available in the country is significant. However, there is scarcity of published report about study on Mx gene in Bangladesh. In an initial study on Mx gene and its diversity in chicken we showed three types Mx gene allele biz. homozygous resistant (R/R), heterozygous (R/S) and homozygous sensitive (S/S) is existed in the sampled chicken population (Jahangir et al., 2015).
2. Materials and Methods
2.1. Blood Sample Collection
A total of 60 blood samples were collected from ten groups (5 from each group) of chicken (Gallus gallus) and duck (Anas platyrhynchos) (Table 1). Of these 50 samples were obtained from Bangladesh Livestock Research Institute (BLRI), Savar, Dhaka, and 10 samples from Nageswary, Kurigram (n=5) and Natore Sadar, Natore (n=5). Chicken groups include Rhode Island Red (RIR), White Leg Horn (WLH), Barred Plymouth Rock (BPR), White Rock (WR), Necked Neck (NN) and Hilly (Hy) and the duck types include Pekin (Pk) Rupali (Rp), Nageshwary (Ny) Commom deshi (CD). About 4% triglyceride was used as an anticoagulant to collect blood. After collection blood was kept in ice box having ice pack and transferred to the laboratory of Animal Biotechnology Division, National Institute of Biotechnology (NIB) and stored at -20oC until use. Laboratory analysis was done at NIB.
2.2. Preparation of Genomic DNA
The DNA was extracted from blood samples using the Phenol: chloroform: iso-amylalcohol method. The extraction procedure was as follows: 200µl of blood was taken in a 1.5 ml micro centrifuge tube and 20µl of proteinase K was added and mixed well. To this, equal volume of Phenol: Chloroform: Iso-amyl alcohol (25:24:1) was added, then gently mixed and centrifuged at 8000Xg for 2 minutes under refrigerated condition. The upper aqueous phase was transferred to a new micro centrifuge tube and above procedure was repeated. The upper aqueous phase was transferred to a new micro centrifuge tube and equal volume of chloroform was added, then gently mixed and centrifuged at 8000Xg for 2 minutes under refrigerated condition. Three molar sodium acetate was added to one tenth of the volume of protein free DNA solution (upper aqueous phase) and two volumes of absolute ethanol was added and kept at –80oC for one hour followed by centrifugation at 15000Xg for 15 minutes. The DNA pellet was re-suspend in 70% ethanol and centrifuged at 15000Xg for 15 minutes and the DNA pellet was dried till no more ethanol left in the tube. DNA was suspended in 50μl of nuclease free water and stored at -20oC.
2.3. Primers
Two sets of primers were used in this study. One set designed by Sironi et. al., (2010) was used to detect the Mx gene. These primers could amplify ~299bp fragments of Mx gene. Another set of mismatched primers designed by Seyama et al., (2006) were used to amplify 100bp fragments of Mx gene followed by restriction enzyme analysis.
2.4. Polymerase Chain Reaction (PCR)
The PCR reaction mixture consisted of Genei red dye Master mix with enzyme12.5μl, forward and reverse primer each 1μl (20 pmole); template DNA 2μl and water 8.5μl. The cycling profile comprised an initial denaturing step for 5 min at 94oC, followed by 35 cycles of 94oC for 1 min, 60oC for 1 min, and 72oC for 1 min, and a final extension at 72oC for 5 min (Seyama et al., 2006). For other primer specified conditions were used (Sironi et al., 2010). Amplicons were analyzed by gel electrophoresis in a 2% agarose gel using TAE buffer and stained with ethidium bromide.
2.5. Restriction Fragment Length Polymorphism (RFLP)
The PCR products obtained using NE-F2 and NE-R2/R primers were digested with the restriction endonuclease RsaI (Biolabs, USA) and those obtained using NE-F2 and NE-R2/S primers were digested with SspI (BioLabs, USA). The digestion reaction (10µl) consisted of nuclease free water 3µl, compatible 10X buffer 1µl, specific restriction enzyme 1µl and PCR product 5µl. The reaction mixture was incubated at 37oC water bath for 1hour. Upon digestion the products were electrophoresed in 3% agarose gel containing ethidium bromide. DNA was visualized with transilluminator.
2.6. Sequencing and Phylogenetic Analysis
PRC product was purified using EZ-10 Spin Column PCR Products Purification Kits-BS363 (Bio Basic Inc, Canada) according to manufacturer’s instruction and stored at -20oC. Purified PCR product was sequenced by dideoxy chain terminating method using Dye Terminator Cycle Sequencing FS Ready Reaction Kit (Applied Biosystems, USA). The nucleotide sequencing reaction mixture contained 0.5μl purified PCR product, 2μl of primer (4 pmol/μl), 4μl of BigDye Terminator v3.1 (Cycle Sequencing Kit; ABI, Foster City, CA), and 4μl of BigDye® Terminator v1.1, v3.1 5X sequencing buffer (Applied Biosystems), and double distilled water amount necessary to make final volume of 20μl, according to the manufacturer’s instructions. The sequence reaction was performed as follows: denaturation for 1 min at 94°C, followed by 25 cycles of PCR amplification, with each cycle consisting of 45 sec of denaturation at 96°C, 30 sec of annealing at 50°C and 4 min of elongation at 60°C. Then the product was purified by BigDye® XTerminator purification kit (Applied Biosystems, USA). The primer used forPCR was used in sequencing reaction. Sequencing was done from both forward and reverse direction. Nucleotide sequences were determined using an automated DNA sequencer 3110 Genetic Analyzer (Applied Biosystems, USA) and edited with SeqEscape V2.6 software. Multiple sequence alignments and processing were performed with the Molecular Evolutionary Genetics Analysis (MEGA) version 4.1.0 software (Tamura et al., 2007) with an engine based on the ClustalW algorithm. Blast searches were used to retrieve the homologous sequences from the GenBank database. The phylogenetic analyses were performed using neighbor joining tree inference analysis.
3. Results
3.1. Amplification of ~299bp Mx Gene Fragment
A total of 55 DNA samples of chicken and duck were subjected to PCR with primer reported by Sironi et al., (2010) and about ~299bp fragment (Fig. 1) of Mx gene was found in only 24 (43.6%) samples (Table 1). The overall detection of Mx gene by this primer set was 43.64%. Detection rate was found to ranges from 60.0-100.0% in chicken and 0-40% in duck. However, average detection rate was 77.8 and 10.7% in chicken and duck, respectively.
Table 1. Detection of Mx gene in chicken and duck.
| Species & Source | Group of chicken and duck | No. of sample collected | No. of sample used in PCR | No. of sample positive | % positive |
| Chicken | |||||
| BLRI | White Leg Horn | 5 | 5 | 4 | 80 |
| White Rock | 5 | 5 | 4 | 80 | |
| Rhode Island Red | 5 | 5 | 4 | 80 | |
| Barred Plymouth Rock | 5 | 3 | 2 | 66.6 | |
| Hilly | 5 | 5 | 3 | 60 | |
| Necked neck | 5 | 4 | 4 | 100 | |
| Total | 30 | 27 | 21 | 77.8 | |
| Duck | |||||
| BLRI | Pekin | 5 | 5 | 0 | 0 |
| Rupali | 5 | 5 | 0 | 0 | |
| Nageshwari | 5 | 5 | 2 | 40 | |
| Common deshi | 5 | 3 | 0 | 0 | |
| Others | Nageshwari (Kurigram) | 5 | 5 | 1 | 20 |
| Common deshi ( Natore) | 5 | 5 | 0 | 0 | |
| Total | 30 | 28 | 3 | 10.7 | |
| Overall | 60 | 55 | 43.6 |
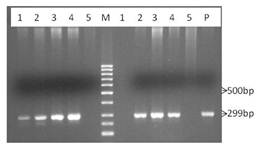
Fig. 1. Amplification of 299bp fragments of Mx gene from chicken and duck by Sironi’s primer, Lane 1-5: Amplification of Mx gene from chicken (left) and duck (right), Lane M: Marker; Lane P: Positive control
3.2. Amplificationof 100bp Fragment of Mx Gene by Mismatched Primers
A total of 37 samples (27 from chicken and 10 from duck) were tested by PCR with mismatched primers NE-F2, NE-R2/R and NE-R2/Sreported by Seyama et al., (2006). Overall 75.7% (28/37) samples were found positive (Table 2). The PCR products obtained using the mismatched primers were 100bp fragment of Mx gene (Fig. 2). Detection rate in chicken with mismatched primers was 40.0-100.0%. On the other hand in duck, the detection rate with NE-F2, NE-R2/R primerwas 100% while with NE-F2, NE-R2/S it was 66.6-100.0%.
Table 2. Amplification of 100bp Mx gene fragment from chicken and duck using mismatched primers.
| Species & source | Group of chicken and duck | No. of sample tested by PCR | PCR result with NE-F2, NE-R2/R primers | PCR result with NE-F2, NE-R2/S primers | ||
| Positive | % positive | Positive | % positive | |||
| Chicken | ||||||
| BLRI | White Leg Horn | 5 | 3 | 60.0 | 4 | 80.0 |
| White Rock | 5 | 4 | 80.0 | 4 | 80.0 | |
| Rhode Island Red | 5 | 3 | 60.0 | 3 | 60.0 | |
| Barred Plymouth Rock | 2 | 2 | 100.0 | 2 | 100.0 | |
| Hilly | 5 | 4 | 80.0 | 4 | 80.0 | |
| Necked neck | 5 | 2 | 40.0 | 2 | 40.0 | |
| Total | 27 | 18 | 66.7 | 19 | 70.4 | |
| Duck | ||||||
| BLRI | Pekin | 2 | 2 | 100.0 | 2 | 100.0 |
| Rupali | 3 | 3 | 100.0 | 3 | 100.0 | |
| Nageshwary | 2 | 2 | 100.0 | 2 | 100.0 | |
| Common Deshi | 3 | 3 | 100.0 | 2 | 66.7 | |
| Total | 10 | 10 | 100.0 | 9 | 90.0 | |
| Overall | 37 | 28 | 75.7 | 28 | 75.7 | |
3.3. Diversity of Mx Gene in Chicken and Duck
A total of 27 PCR positive samples (18 from chicken and 9 from duck) with mismatched primers were used to determine the genetic variation/diversity of Mx gene in chicken and duck. PCR products obtained with NE-F2 & NE-R2/R primer were digested with restriction enzyme RsaI while PCR products obtained with NE-F2 and NE-R2/S were digested with SspI. Both the enzymes would cut the 100bp PCR product into two 73 and 27bp length fragments. Three types of results viz. complete, partial or no digestion were observed with each of the restriction enzyme. Complete, partial and no digestion with RsaI indicate sensitive (homozygous denoted as S/S) variable (heterozygous denoted as R/S) and resistant (homozygous denoted as R/R), to influenza, respectively. While complete, partial and no digestion with SspI indicate resistant (homozygous denoted as R/R), variable (heterozygous denoted as R/S) and sensitive (homozygous denoted as S/S) to influenza, respectively. Interpretation of restriction enzyme analysis of 27 samples with both enzymes is presented in Table 3. Of 18 samples of chicken 38.9% (7/18) were found homozygous resistant, 27.0% (5/18) heterozygous and 33.3% (6/18) homozygous sensitive. On the other hand 11.1, 22.2 and 66.6% samples in duck were found homozygous resistant, heterozygous and homozygous sensitive, respectively. Comparatively ducks were found more sensitive.
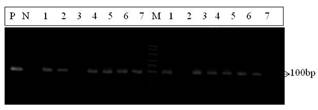
Fig. 2. Amplification of 100bp fragments of Mx gene by PCR from chicken. Lane 1-7 (left): Amplification by primer NE-F2 & NE-R2/R, Lane 1-7 (right): Amplification by primer NE-F2 and NE-R2/S, Lane M: Marker, Lane P: Positive control; Lane N: Negative control
Table 3. Diversity of Mx gene in chicken and duck
| Species | Group of chicken and duck | No. of sample tested | Diversity of Mx gene | ||
| R/R | R/S | S/S | |||
| Chicken | |||||
| BLRI | White Leg Horn | 3 | 1 | 1 | 1 |
| White Rock | 4 | 1 | 1 | 2 | |
| Rhode Island Red | 3 | 0 | 1 | 2 | |
| Barred Plymouth Rock | 2 | 1 | 0 | 1 | |
| Necked Neck | 4 | 3 | 1 | 0 | |
| Hilly | 2 | 1 | 1 | 0 | |
| Total | 18 | 7 (38.9%) | 5 (27.0%) | 6 (33.3%) | |
| Duck | |||||
| BLRI | Pekin | 2 | 0 | 0 | 2 |
| Rupali | 3 | 0 | 1 | 2 | |
| Nageshwary | 2 | 0 | 0 | 2 | |
| Common Deshi | 2 | 1 | 1 | 0 | |
| Total | 9 | 1 (11.1%) | 2 (22.2%) | 6 (66.7%) | |
| Overall | 27 | 8 (29.6%) | 7 (25.9%) | 12 (44.4%) | |
R/R: Homozygous resistant, R/S: Heterozygous, S/S: Homozygous sensitive.
3.4. Phylogenetic Analysis
One of the PCR products of native chicken necked neck was sequenced. Obtained 98bp sequence has 95% homology with the sequences of Gallus gallus located on the right side of the middle line drawn for necked neck (Fig.3). These sequences were derived from Gallus gallus clone EL494Mx1 (Accession No. 308212361), Gallus gallus Mx protein (Mx) gene (Accesssion No. 257219904), Gallus gallus isolate While Leghorn Mx protein (Mx) mRNA (Accession No. 146744131), Gallus gallus isolate WL_42Mx protein (Mx) mRNA (Accession No. 146744165), Gallus gallus isolate 25 breed White Leghorn MX (MX) gene (Accession No. 111182889) and Gallus gallusMx protein mRNA (Accession No. 164551495). Sequences located left side of the middle line has 93% homology. They also derived from Gallus gallus Mx protein mRNA. Accession number of each sequence is depicted in the tree.
4. Discussion
Chickens are natural hosts to influenza virus (Easterday, 1975) and many other viruses, and virus infection causes serious illness or death in chickens. The avian influenza virus (AIV) is also infectious to humans and has led to an acute condition in some cases (Subbarao et al., 1998). Selective breeding of AIV-resistant chickens would be beneficial for both the livestock industry and human health. Therefore present study was conducted to determine the avian influenza resistant gene or Myxovirus resistant gene (Mx) and its diversity in chicken and duck. For this purpose blood samples from six groups of chicken and four groups of duck were analyzed by PCR and PCR-RFLP. In this study two different sets of primers were used to detect Mx gene and its diversity in the sampled population. These primers are reported by Sironi et al., (2010) and Seyama et al., (2006). We used primer set reported by Sironi et al.,(2010) to amplify ~299bp fragment only for detection of Mx gene. On the other hand Mx gene detection and diversity analysis was done by PCR-RFLP with mismatched primers and restriction enzymes reported by Seyama et al., (2006). We could amplify the specific sized DNA using these reported primers (Fig. 1 and 2). However, the detection rate varies with the different sets of primers (Table 1 and 2). The overall Mx gene detection rate was higher (75.7%) with the primer reported by Seyamaet al., (2006) than the primer reported by Sironi et al., (2010) (43.6%) (Table 1 and 2). These variations might be due to different primers because the primers were designed based on different location of Mx gene with different targets. It is reported that Sironi-F primer anneals to the last intron of the Mx gene and Mx-Sironi-R primer anneals to the last axon on the gene (Sironi et al., 2010) while NE2-F2 bind to axon 14 of the Mx gene Seyama et al., (2006). Samples with negative result were tested at least three times to minimize error to confirm the true negativity. From these findings it seems primer reported by Seyama et al., (2006) would be more suitable to determine the Mx gene in chicken and duck.
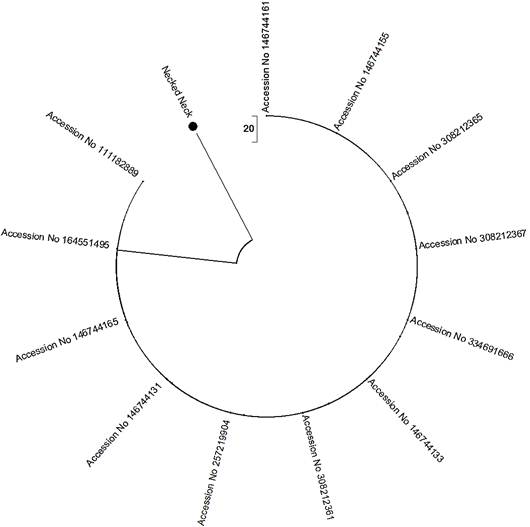
Fig. 3. Phylogenetic analysis of portion (98bp) of Mx gene of indigenous chicken necked neck. An unrooted neighbor joining tree of nucleotide sequences of Mx gene were generated. Sequences right and left side of the midline has 95 and 93% homology, respectively with the sequence of the Mx gene of necked neck (marked in the three with black circle) in the middle of the tree.
Mx protein induced by type I IFN is known to inhibit the multiplication of various viruses, including influenza virus (Lee and Vidal, 2002). However, chicken Mx protein in German White Leghorn lacked antiviral activity for both influenza and vesicular stomatitis viruses (VSV) (Bernasconi et al., 1995). Similarly, Benfield et al., (2008) reported that breed Shamo, which has an Asparagine at amino acid 631 is void of activity against the H5N1. These variations are due to variation in Mx gene. Because it is reported that a specific amino acid substitution between Asn and Ser at position 631 determines the differential antiviral activity of chicken Mx protein (Ko et al., 2002). In the present study we determine the diversity of Mx gene and we found diversity is existed in the tested samples. Three types of results R/R, R/S and S/S were found. Diversity of Mx gene not only observed among the groups but also within the groups (Table 3). Indigenous chicken (Hilly and Necked neck) were found either resistant or heterozygous. We have tested only six samples and none was grouped into sensitive type. Our findings comply with the findings of Seyama et al. (2006). They investigated 271 DNA of commercial and indigenous chickens from different sources and found resistant, sensitive and heterozygous Mx gene allele within and between the groups of chicken. Similar results were also reported by Sartika et al. (2011), Sulandari et al. (2009). They also reported that indigenous chicken has more frequency of resistant type Mx gene allele. Findings of the present study regarding distribution of Mx gene have likeness with the findings of Seyama et al., (2006). We found 29.6, 44.4 and 25.9% of the tested sample asresistant, sensitive and heterozygous Mxgene alleleic, respectively. While Seyama et al. (2006) reported 33.95% resistant, 52.40% sensitive and 13.65% heterozygous allele in the tested samples (n=271). Present finding also has likeliness with the findings of our previous work where we found 42.86% homozygous resistant (R/R), 42.86% homozygous sensitive (S/S) and 14.29% heterozygous.
Outbreak of HPAI was first occurred in Bangladesh in 2007 and since then it is continued to occur across the country. The disease is mostly occurred in commercial chicken (n=499) than native chicken (n=57) (Giasuddin et al., 2013). Though duck are considered to be resistant, an HPAI outbreak in native ducks in Netrokona district was reported in 2011 (ICDDR,b, 2013). However, no such occurrences of HPAI were reported from rest part of the country. But in Indonesia ducks are appeared to be mostly affected species with 25-50% mortality in juvenile, 19% in backyard and 33% in commercial duck (FAO, 2014). These findings suggest there may be some host factor responsible for lower occurrences of HPAI in native poultry and duck population in Bangladesh. Presence of Mx gene in the chicken and duck population may be that host factor. We have examined only 9 duck samples for Mx gene diversity analysis in this study and found that 66.6% samples were belonging to sensitive group while only 11.1% belong to resistant type and rest 22.2 heterozygous. This finding contradicts with the report of lower number of HPAI occurrences in our duck. Because if ducks are more sensitive then more outbreak should be occurred in this group and eventually reported number should be more in duck. However, our findings comply with the notion that ducks are reservoir of AIV. For harbouring and maintenance virus should undergo replication cycle in the host cell. But if the duck’s cytoplasmic Mx protein inhibits the replication then AIV could not maintain in duck.
5. Conclusion
The Mx gene is amplified from all the six groups (two indigenous & four exotic breeds) of chicken and four groups of duck. Three types of Mx gene allele are found in the study. These are homogenous resistant ((R/R), homogenous sensitive (S/S) and heterozygous (R/S). Diversity is found within and between groups of chicken and duck. Sequence analysis of portion of Mx gene of necked neck indigenous chicken showed 95% homology with Mx gene from White Leghorn and other chicken Mxgene.
References
- Abdelwhab, E. M., and H.M. Hafez. 2012. Insight into alternative approaches for control of avian influenza in poultry, with emphasis on highly pathogenic H5N1. Viruses, 4: 3179-3208.
- Aebi, M., J. Fah, N. Hurt, C.E. Samuel, D. Thomis, L. Bazzigher, J. Pavlovic, O. Haller and P. Staeheli. 1989. cDNA structures and regulation of two interferon-induced human Mx proteins. Mol. Cell. Biol.,9:5062-5072.
- Baise, E., G.G. Pire, M.I. Leroy, J.I. Garardin, N. Goris, K.D. Clercq, P. Kerkhofs and D. Desmecht. 2004. Conditional expression of type I interferon-induced bovine Mx1GTPase in a stable transgenic vero cell line interferes with replication of vesicular stomatitis virus. J. of Interferon & Cytokine Res., 24:513-521.
- Bazzigher, L., A. Schwarz and P. Staeheli. 1993. No enhanced influenza virus resistance of murine and avian cells expressing cloned duck Mx protein. Virol., 195:100-112.
- Benfield, C.T.O., J.W. Lyall, G. Kochs and L.S. Tiley. 2008. Asparagine 631 variants of the chicken Mx protein do not inhibit influenza virus replication in primary chicken embryo fibroblasts or in vitro surrogate assays. J. Virol., 82:7533-7539.
- Bernasconi, D., U. Schultz and P. Staeheli. 1995. The interferon-induced Mx protein of chickens lacks antiviral activity. J. of Interferon & Cytokine Res., 15: 47-53.
- Bhuiyan, A., M.S.A. Bhuiyan and G.K. Deb. 2005. Indigenous chicken genetic resources in Bangladesh: Current status and future outlook. Anim. Genet. Resources Informatio., 36: 73-84.
- Bhuiyan, M.S.A., S. Chen, S. Faruque A. Bhuiyan and A. Beja-Pereira. 2013. Genetic diversity and maternal origin of Bangladeshi chicken. Mol.Boil. Rep,40: 4123-4128.
- FAO.2014. Update on the continuous spread and expansion of H5N1 highly pathogenic avian influenza: Clade 2.3.2.1 in Asia (2010–2012), FOCUS on, No. 7, January 2014. Rome.
- Garber, E.A., C.I. Rosenblum, H.T. Chute, L.M. Scheidel and H. Chen. 1991. Avian retroviral expression of luciferase. Virol., 185: 652-660.
- Giasuddin, M., M.E. Haque, A.H.M. Kamal, M.R. Islam, A. Jahangir, E.H. Chowdhury, M.J.F.A. Taimur andM.H. Rahman. 2013. Outbreak evaluation of highly pathogenic avian influenza in Bangladesh. Bang. J. Lives. Res. 19 (1-2): 41-45
- Haller, O., M. Frese, D. Rost, P.A. Nuttall and G. Kochs. 1995. Tick-borne thogoto virus infection in mice is inhibited by the Orthomyxovirus resistance gene product Mx1.J Virol., 69:2596-2601.
- Haller, O., S. Stertz and G. Kochs. 2007. The MxGTPase family of interferon-induced antiviral proteins. Microbes and Infection, 9:1636-1643.
- Horisberger, M.A. 1995. Interferons, Mx genes, and resistance to influenza virus. American J. Respir. Crit. care Med., 152:S67.
- ICDDR,b. 2013. Unusual waterfowl mortality due to highly pathogenic avian influenza A (H5N1) virus in Netrokona, Bangladesh 2011. Health and Science Bulletin.11 (1). March 2013.
- Jahangir, A., H. Mahmudul, H. Kamrul, M. Giasuddin, M.R. Hafizur, A. Shahin and M.J.F.A. Taimur. 2015. Diversity of Myxovirus resistance gene (Mx gene) in chicken. Proceedings of the 9th International Poultry show and Seminar of World’s Poultry Science Association- Bangladesh Branch held on 19-21 February 2015. pp. 165-172
- Janzen, C., G. Kochs and O. Haller. 2000. A monomeric GTPase-negative MxA mutant with antiviral activity. J. Virol.,74:8202-8206.
- Jin, H.K., A. Takada, Y. Kon, O. Haller and T. Watanabe. 1999. Identification of the murine Mx2 gene: interferon-induced expression of the Mx2 protein from the feral mouse gene confers resistance to vesicular stomatitis virus. J. Virol.,73:4925-4930.
- Ko, J.H., H.K. Jin, A. Asano, A. Takada, A. Ninomiya, H. Kida, H. Hokiyama, M. Ohara, M. Tsuzuki and M. Nishibori. 2002. Polymorphisms and the differential antiviral activity of the chicken Mx gene. Genome Res,12:595-601.
- Landis, H., A. Simon-Jadicke, A. Klati, C. Di Paolo, J.J.R. Schnorr, S. Schneider-Schaulies, H. P. Hefti and J. Pavlovic. 1998. Human MxA protein confers resistance to Semliki Forest virus and inhibits the amplification of a Semliki Forest virus-based replicon in the absence of viral structural proteins. J. Virol.,72:1516-1522.
- Lee, S.H. and S.M. Vidal. 2002. Functional diversity of Mx proteins: variations on a theme of host resistance to infection. Genome Res.,12:527-530.
- Li, X.Y., L.J. Qu, Z.C. Hou, J.F. Yao, G.Y. Xu and N. Yang. 2007. Genomic structure and diversity of the chicken Mx gene. Poultry Scienc., 86:786-789.
- Livant, E.J., S. Avendano, S. McLeod, X. Ye, S.J. Lamont, J.C.M. Dekkers and S.J. Ewald. 2007. MX1 exon 13 polymorphisms in broiler breeder chickens and associations with commercial traits. Anim. Genet., 38: 177-179
- Meier, E., G. Kunz, O. Haller and H. Arnheiter. 1990. Activity of rat Mx proteins against a rhabdovirus. J. Virol.,64:6263-6269.
- Muller, U., U. Steinhoff, L.F. Reis, S. Hemmi, J. Pavlovic, R.M. Zinkernagel, and M. Aguet. 1994. Functional role of type I and type II interferons in antiviral defence. Sci.,264:1918-1921.
- Pavlovic, J., O. Haller and P. Staeheli. 1992. Human and mouse Mx proteins inhibit different steps of the influenza virus multiplication cycle. J. Virol.,66:2564-2569.
- Rothman, J.H., C.K. Raymond, T. Gilbert, P.J. O'Hara and T.H. Stevens. 1990. A putative GTP binding protein homologous to interferon-inducible Mx proteins performs an essential function in yeast protein sorting. Cell, 61:1063-1074.
- Sartika, T., S. Sulandari, and M.S.A. Zein. 2011. Selection of Mx gene genotype as genetic marker for Avian Influenza resistance in Indonesian native chicken. Pp. S37. BMC proceedings. BioMed Central Ltd.
- Seyama, T., J.H. Ko, M. Ohe, N. Sasaoka, A. Okada, H. Gomi, A. Yoneda, J. Ueda, M. Nishibori and S. Okamoto. 2006. Population research of genetic polymorphism at amino acid position 631 in chicken Mx protein with differential antiviral activity. Biochem. Genet,44:432-443.
- Sironi, L., P. Ramelli, J.L. Williams and P. Mariani. 2010. PCR-RFLP genotyping protocol for chicken Mx gene G/A polymorphism associated with the S631N mutation. Genet. Mol. Res.,9:1104-1108.
- Subbarao, K., A. Klimov, J. Katz, H. Regnery, W. Lim, H. Hall, M. Perdue, D. Swayne, C. Bender, J. Huang, M. Hemphill, T. Rowe, M. Shaw, X. Xu, K. Fukuda and N. Cox. 1998. Characterization of an avian influenza A (H5N1) virus isolated from a child with a fatal respiratory illness. Science. 279, 393–396.
- Sulandari, S., M. SyamsulArifinZein, D. Astuti and T. Sartika. 2009. Genetic polymorphisms of the chicken antiviral Mx gene in a variety of Indonesian indigenous chicken breeds. J. Veteriner., 10: 50-56.
- Schneider-Schaulies, S., J.R. Schneider-Schaulies, A. Schuster, M. Bayer, J. Pavlovic and V. TerMeulen. 1994. Cell type-specific MxA-mediated inhibition of measles virus transcription in human brain cells. J. Virol., 68:6910-6917.
- Schwemmle, M., K.C. Weining, M.F. Richter, B. Schumacher and P. Staeheli. 1995. Vesicular stomatitis virus transcription inhibited by purified MxA protein. Virol., 206:545-554.
- Zurcher, T., J. Pavlovic and P. Staeheli. 1992. Mechanism of human MxA protein action: variants with changed antiviral properties. Embo J.,11(4):1657-61.



