Rapid Diagnosis of Acute Respiratory Infections by Multiplex Endpoint PCR Technology
Aurelian Udristioiu1, *, Manole Cojocaru2, Dana Alexandra Maria Panait2, Nica Badea Delia3
1Emergency County Hospital Targu Jiu & UCB University, Clinical Laboratory, City Targu Jiu, Romania
2TituMaiorescu University, Faculty of Medicine, Physiology Department, City Bucharest, Romania
3Constantin Brancusi University, Faculty of Medical Science and Behaviors, City Targu Jiu, Romania
Abstract
Introduction: PCR offer a number of potential advantages, results are available in a matter of hours rather than days, the extreme sensibility facilitates detection of even minutes the amounts of pathogen DNA in clinical samples and the test is not significantly affected by prior administration of antibiotics. Aim: The aim of this work was to rapidly identify the antibiotic resistance the monitoring of pathogen grow that the patients admitted in Hospitalization Intensive Care Unit of Emergency County Hospital Targu Jiu starting in December/2013. Method: The Analyzer Unyvero™ Pneumonia Application was used in detection of pneumonia associated pathogens and their antibiotic resistance genes using the Unyvero™ System following PCR, pathogen species with sequencing of the amplified microbial DNA. Results: The main pathogens of community acquired pneumonia are Streptococcus pneumonia (5 cases), and Klebsiella pneumonia (4cases), other important agents are "atypical", such as Haemophilus Influenzae, Chlamidophila pneumonie and Moraxela cataralis. A case with Acinetobacter baumani and Proteus Sp.are also widely resistance to mefA gene / ermB gene as all cases of analyzed. The more frequency of gene resistant (9 cases) are ermA gene / ermC / ermB for Staphilococcus aureus and the gene tem+shv / gene / ctx-M with the Chromosomal mutation (4 cases), as gyrA83_87 Ecoli / Pseu for Klebsiella pneumonia agents. Also most resistance antibiotics was Makrolides (9 cases and Lincosamides (4 cases) and this cases have chromosomial integrates. The most resistance microbe, Pseudomonas aeruginosa (1 case), has registered multi drugs resistance [MDR]*.Conclusions: The Unyvero™ results were available 2 days before the primary microbiology report and 3 days before the final confirmation results, obtained by microbiology culture. The Unyvero Analyzer only provides rapid data to support the therapeutic decision of currant medic.
Keywords
Antibiotic Resistance, PCR, Gene, DNA, Resistance Markers, Microbiology Report
Received: December 14, 2014
Accepted: January 8, 2015
Published online: February 10, 2015
@ 2015 The Authors. Published by American Institute of Science. This Open Access article is under the CC BY-NC license. http://creativecommons.org/licenses/by-nc/4.0/
1. Introduction
While microbiological culture is likely to remain a gold standard for infection diagnosis, there is growing interest at the potential of PCR technology to provide early, time critical information based on detection and recognition of bacterial or fungal pathogen DNA.
Following PCR, pathogen species present can be identified sequencing of the amplified DNA. The assay for detection and identification of a defined panel of 25 bacterial of 25 bacterial and fungal pathogen known to cause a majority of acute or chronic respiratory infections.
The need of sophisticated arsenal of new technological tools in the microbiology lab is dictated by two recent events. The first is the emergence and growth of though strain of microbe resistant to antibiotics, even the new antibiotics is drying up (ex. Microorganism such as K. pneumonia and E. coli have become resistant to third generation cephalosporins as well as carbapenems. The focus of researches now is to move beyond detecting single analytes to multiplex targets and detect more pathogens from a single specimen, ex. Sputum, (1).
1.1. Principle of the Analysis
The Unyvero™ Pneumonia Application automates has integrates in a disposable cartridge, genomic DNA purification, eight parallel multiplex end-point PCR reactions and the qualitative detection of the target amplicons after hybridization onto an array, [Photo 1].
Photo 1. AnalyzerUnyvero™ Pneumonia
1.2. Technique
1.The patient sample is pipetted into the Unyvero™ Sample Tube using the Unyvero™ Sample.
2.Transfer Tool, closed with the Unyvero™ Sample Tube Cap, and lysed with the Unyvero™ Lysator.
3.Subsequently, the Unyvero™ Sample Tube and the Unyvero™ Master Mix Tube are inserted into the Unyvero™ Pneumonia Cartridge.
4.The Unyvero™ Pneumonia Cartridge is then inserted into the Unyvero™ Analyzer, which processes it automatically.
The supplied software guides the user through the entire work flow. A bar code reader allows the entry patients data, checks the shelf-life of consumables and stores their lot numbers. The full analysis should take approximately 30 minutes. The analysis of the patient samples is shown by grey test bars on the overview screen. To view the results, tap on the corresponding blue test bar. A screen opens and shows the following buttons: Summary, Microorganisms, Resistance Markers, information. In the middle of the screen, the respective antibiotic classes for which a therapeutic failure must be considered if they were administered are displayed. On the right side of the screen, the common microbial source of the resistance markers is displayed.
1.3. Detection Limits
Detection limits for each pathogen was determined with pathogen dilutions in buffer. At the concentration of 106 pathogens / mL all analytes are detected with the Unyvero™ P50 Pneumonia Cartridge. In addition the majority of the analytes are positive at a concentration of 104 pathogens / Ml (S. marcescens. S. maltophilia, A. baumannii, L. pneumophila, S. aureus, M. morganii, K. pneumoniae, K. oxytoca, P. aeruginosa). Detection limits for resistance analytes can be determined with DNA fragment dilutions. At a concentration of 105 copies / mL all resistance analytes are detected.
1.4. Interfering Substances
Interferences were tested in suitable pools with respiratory drugs, common antibiotics and sample media or individually for example for lysis buffer, blood, human DNA, and common respiratory pathogens, which might be present in respiratory samples. Worst case concentrations were used according to CSLI guideline "EP7-A2. No interference was observed.
1.5. Sensitivity & Specificity
The Unyvero™ Pneumonia Application achieved an overall sensitivity of 75.5 % (sensitivity per analyte between 50% and 100%, depending on the microorganism) at an overall specificity of 95.2% (72.3% to 100%, depending on the microorganism). For rare pathogens, the number of cases was insufficient to establish sensitivity and specificity data. For detected resistance markers (mefA, ermA, ermB, ermC, tem, shv, dha, oxa51 like, ctxM, mecA, ebc, quinolone resistances in E. coli and P. aeruginosa) in 26 cases out of 32 antibiotic resistant pathogens a correlation between Unyvero P50™ results with the antibiogram was demonstrated. Curetis is currently conducting a prospective European multicenter clinical trial to generate more clinical performance data.
2. Method
Sample type aspirate sputum, at the 11 patients (7 gender males, mean age 65 years, gender female, mean age 55 years), admitted in Hospitalization Intensive Care Unit and The Unyvero™ Pneumonia Application was performed in the day after specimen collection in Department of Microbiology from Clinical Laboratory Analyses of Emergency County Hospital Targu Jiu.
The selection of the samples at the patients admitted in Intensive Care Unit (ICU) with community acquired pneumonia were based on typical clinical signs of severe infection, in evidences of clinician doctors which were included the symptoms such as increased fever, positive X-ray, presence of purulent sputum and on the results of laboratory blood samples with increased white blood cells count( >15000/mm³), VSH (>40 mm/h), Fibrinogen (>450 mg/dl) and Protein C Reactive (>12 mg/dl).
These signs of severe acute infection were primordially for faster results in pneumonia testing. Such quick results from laboratory are perquisite for giving adequate antibiotic treatment as early as possible in order to improve the standard of care.
3. Results
The main pathogens of community acquired pneumonia are Streptococcus pneumonia, (5cases) and Klebsiella pneumonia (4cases), other important agents are "atypical", such as Haemophilus Influenzae, Chlamidophila pneumonie and Moraxela cataralis, [Table 1].
Table 1. Resistance markers of the Pneumonia panel and the resulting antibiotic resistances
| No. IDL | Microorganisms detected | Antibiotic resistance | Gene resistance |
| 1310_1 | Klebsiella pneumonia | Makrolides /], [ermB],].Lincosamides | ermB gene |
| 1310_2 | Streptoccocus Sp. | Makrolides. | ermB gene/tem gene |
| 1310_3 | Staphilococcus aureus Other/Fungi: Chlamidophila pneumonie | Penicilins (tem) | ermB gene/tem gene Chromosomal mutation; Pseud. aeruginosa, (gyrA83- Ecoli |
| 1310_4 | Klebsiella pneumonia | Makrolides / Lincosamides | ermB gene |
| 1310_5 | Proteus Sp. Other/Fungi: Haemophilus Influenzae, Chlamidophila pneumonie | Makrolides Oxacilin | ermB gene/oxa51 Chromosomal mutation; Escherichia Coli (gyrA83-87_Ecoli ). |
| 1311_1 | Streptococcus pneumonia Moraxela cataralis | Makrolides, Oxacillin Lincosamides, | ermB gene/tem gene/ mecA gene |
| 1311_2 | Streptococcus pneumonia | Makrolides / [mefA], [ermB],].Lincosamides Penicilins (tem) | mefA gene / ermB gene / tem |
| 1311_3 | Streptococcus pneumonia Pseudomonas aeruginosa | Makrolides / [mefA], [ermB],].Lincosamides [MDR]* [int1]. | mefA gene / ermB gene [int1gene]. |
| 1311_4 | Klebsiella pneumonia | Penicilins (shv) | mefA gene / shv gene [int1gene] /sul 1 gene Chromosomal mutation; (gyrA83_3Pseu). |
| 1311_5 | Streptococcus pneumonia Acinetobacter baumani | Makrolides / [mefA], [ermB],].Lincosamides | mefA gene / ermB gene |
| 1312_1 | Staphilococcus aureus Klebsiella pneumonia | Makrolides / [mefA], [ermB],].Lincosamides 3rd Gen Cephalosporins [tem+shv], [ctx-M]. | ermA gene / ermC / ermB tem+shv / gene / ctx-M g Chromosomal mutation; gyrA83_87 Ecoli / Pseu |
*Multi drugs resistance
A case with Acinetobacter baumani and Proteus Sp.are also widely resistance to mefA gene / ermB gene as all cases of analyzed. The more frequency of gene resistant (9 cases) are ermA gene / ermC / ermB for Staphilococcus aureus and the gene tem+shv / gene / ctx-M with the Chromosomal mutation (4 cases), as gyrA83_87 Ecoli / Pseu for Klebsiella pneumonia agents.
Also most resistance antibiotics was Makrolides (9 cases and Lincosamides (4 cases) and this cases have chromosomial integrates {Fig1]. The β-lactam antibiotics work by inhibiting the cell wall synthesis by binding to so-called penicillin-binding proteins (PBPs) in bacteria and interfering with the structural crosslinking of peptidoglycans.
The most resistance microbe, Pseudomonas aeruginosa(1 case), has registered multi drugs resistance[MDR]*.
4. Interpret Results
The green boxes on the Analyzer of The Unyvero™ Pneumonia Application and values do loosely correlate with the amount of detected DNA and therefore with the number of pathogens in a given patient sample – however, the number of pathogens obtained by culture does not always correlate with the number of pathogens in a sample due to limitations of growth.
The numbers next to the green boxes are artificially created and normalized numbers without a measurable unit. These numbers are reflecting a threshold value depending on the species and serve to give an aid to quantification. (< 250 no green box; 250 – 499 one green box, 500 – 999 twogreen boxes, >= 1000 three green boxes). It was a specific customer demand to have some form of number, [Photo 2].
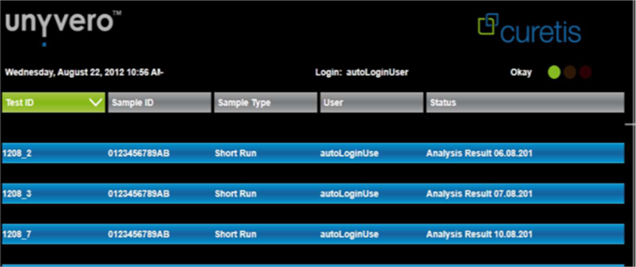
Photo 2. Analysis of the patient samples shown by grey test bars on the overview screen.
![]() Detected,
Detected, ![]() Invalid measurement,
Invalid measurement, ![]() Not detected
Not detected
However, a clinician may still take this data into consideration e.g. in an immune compromised patient and with certain pathogens. Still, a clinician may take this data into consideration e.g. in an immune compromised patient and with certain pathogens. These results were confirmed by microbiology culture; however, the final microbiology result was available 3 days after the Unyvero™ result only. Gram-Stain: Microscopy exam:
The Classical Method Kirby Bauer Disc Diffusion Method
In Culture at 1310_1 raised Klebsiellap neumoniae (+++), Sensible Antibiotics, Amoxicillin, Sulfonamide Cotrimoxazol, Quinolone Moxifloxacin, Monobactame Aztreonam, Chephalosporin Cefotaxim AND Ampicillin R, Amoxic./Clavulanacid R, Piperacillin R, Piperac/Tazobactam R, resistance spread especially on our intensive care unit.
5. Discussions
The Unyvero™ Pneumonia Cartridge can detect the following microorganisms: Acinetobacterbaumannii, Chlamydophila pneumoniae, Enterobacter sp., Escherichia coli, Haemophilus influenzae, Klebsiellap neumoniae, Klebsiella oxytoca, Legionella pneumophila, Moraxella catarrhalis, Morganella morganii, Pneumocystis jirovecii, Proteus sp., Pseudomonas aeruginosa, Serratia marcescens, Staphylococcus aureus, Stenotrophomonas maltophilia und Streptococcus pneumoniae.
Simultaneously, the following genes associated with antibiotic resistance are detected in the same cartridge: ctx-M, (Cephalosporines, Penicillines), dha (Cephalosporines), ebc (3rd Gen. Cephalosporines), ermA(Makrolides / Lincosamides), ermB (Makrolides / Lincosamides) , ermC (Makrolides / Lincosamides) , gyrA83, (E. coli ), gyrA87, int1, kpc, mecA, mefA/E, msrA, oxa51 like, parC, shv, sul1, and tem (Fluoroquinolones, E. coli), (3).
5.1. Transferable Antibiotics Resistance (Resistance Genes)
Most resistance markers that are detected by the Unyvero™ Pneumonia Application are genes, which are transferred by mobile genetic elements like plasmids or integrons 6. Presence of such a gene correlates with a resistance against a particular antibiotic class7. Gene transfer is also more likely in environments where bacteria are in close proximity to each other and in relatively high density such as the gut and oral cavity, (5). In order to control the spread of resistance it is important to have an understanding of the molecular biology of the different mobile genetic elements and of the ecology of the environments in which spread is likely, (6).
Bacteria have become resistant to antimicrobials through a number of mechanism; I, Permeability changes in the bacterial cell wall which restricts antimicrobial access to target sites, II, Active efflux of the antibiotic from the microbial cell, III, Enzymatic modification of the antibiotic, IV, Degradation of the antimicrobial agent, V. Acquisition of alternative metabolic pathways to those inhibited by the drug, VI, Modification of antibiotic targets, VII, Overproduction of the target enzyme, (7).
The major encountered aminoglycoside resistance mechanism is the modification of enzymes. These proteins are classified into three major classes according to the type of modification: AAC (acetyltransferases), ANT (nucleotidyl transferases or adenyltrans- ferases). Macrolides have a similar mode of antibacterial action and comparable antibacterial spectra as two other antibiotic classes, i.e., lincosamides and streptogramins B. Consequently, these antibiotics, although chemically distinct, have been clustered together as Macrolide–Lincosamide–StreptograminB(MLS) antibiotics (8).
The β-lactam antibiotics work by inhibiting the cell wall synthesis by binding to so-called penicillin-binding proteins (PBPs) in bacteria and interfering with the structural crosslinking of peptidoglycans and as such preventing terminal trans-peptidation in the bacterial cell wall. At Staphilococcus meticilin resistance (MLS) the resistance is due to the presence of r-RNA methylases, encoded by the "erm "genes. The other two mechanisms efflux pumps and inactivating genes are encoded by ‘msr "and "ere" determinants, respectively (9, 10, 11).
Gene transfer is also more likely in environments where bacteria are in close proximity to each other and in relatively high density such as the gut and oral cavity. In order to control the spread of resistance it is important to have an understanding of the molecular biology of the different mobile genetic elements and of the ecology of the environments in which spread is likely.
The Unyvero™ result was available 2 days before the primary microbiology report and 3 days before the final classical culture method, confirmation test. A more adequate and result guided antibiotic therapy regime with the usage of an ESBL active carbapenem would have been made possible much earlier. In addition, appropriate hygiene measures could have been taken earlier decreasing the risk of antibiotic, (12).
Most resistance in Enterobacteriaceae is associated with acquired resistance genes, often carried by plasmid various encoding B-Lactamases, aminoglycoside modifying enzymes, r-RNA metylases, anti-folate by-pass enzymes or tetracycline efflux pumps. The carbapenen resistance in emerging in Klebsiella pneumonia involves the genes Oxa-48, Pseu, detectable by PCR. A few resistance of Enterobacteriaceae are largely or entirely mutations; the best example is quinolone resistance, which mostly occurs by mutations gyr/A/B/C genes, (13).
These agents, because of their short generation times and, in some cases, high mutation rates, evolve quickly, resulting in evasion of hostmicrobial defenseinter- and intra-host diversification in real time, and acquired resistance to antimicrobial drugs, [14].Also, these infectious diseases can progress quickly, within hours or days, then successful therapeutic intervention can be assured by the specific PCR assays, [15,16]. Understanding the basic concepts of these analysis steps is important in assessing and addressing the informatics needs of a molecular diagnostics laboratory for the rapid communication between clinicians and laboratory results, especially in cases of emergency. Bioinformatics must become an important component in clinical laboratories, maintaining, and interpreting data from molecular genetics testing. Given the rapid adoption of next generation sequencing (NGS), based clinical testing, service providers must develop informatics work flows that adhere to the rigor of clinical laboratory standards, [17].
6. Conclusion
PCR is a key technology in molecular biology and diagnostics that typically amplifies and quantifies specific DNA fragments in about 3-4 hours.
Rapid molecular diagnosis assays has the potential to speed up pathogen and resistance identification, which may enable clinicians to make an early, informed diagnosis in patients with pneumonia.
The Unyvero only provides data to support the therapeutic decision. While microbiological culture is likely to remain a gold standard for infection diagnosis, there is growing interest at the potential of PCR technology to provide early, time critical information based on detection and recognition of bacterial pathogen DNA.

Figure 1. Therapeutic failure for positive microorganism.
References
- Caliendo AM. Multiplex PCR and emerging technologies for the detection of respiratory pathogen.Clin Infect 2011; (4):5326-5330.
- Rulls© Curetis AG, 2013 Unyvero ™ P50 Pneumonia Application Guide 00118 V2.0, Hamburg, Germany, 2013, http://www.curetis.com/ .
- Kugelberg et al. Antimicrobials, Resistance J AntimicrobChemother 2005; 55(1): 1-5
- Yu et al. Antimicrobials, Resistance J ClinMicrobiol 2004; 42(9): 1-4
- VanHoek A, Mevius D, Guerra B, Mullany P and al. Acquired antibiotic resistance genes a new review article published 2011; 1: 1-28.
- Depardieu A et al. Frontiers in Microbiology. Antimicrobials, Resistance and Chemotherapy. 2011;203: 1-13.
- Wright G.D. Bacterial resistant and antibiotics:enzymaticdegradationandmodification. Adv. DrugDeliv.Rev 2005; 57: 1451–1470.
- Roberts M.C. Update on macrolide-lincosamide, treptogramin, ketolideandoxazolidinone resistance genes, -ermA, ermB, ermCMakrolides / Lincosamides, gyrA83 of E. coli. FEMS Microbiol.Lett 2008; 282: 147–159.
- Bonnet et al. ctx-M 3rdGen, Cephalosporines, Penicillines, Antimicrob Agents. Chemother 2001; 45(8): 1-8.
- Pérez-Pérez et al. dha 3rdGen. Cephalosporines, J ClinMicrobiol 2002; 40(6): 1-6.
- Pérez-Pérez et al. ebc 3rd Gen. Cephalosporines Pérez-Pérez et al, J ClinMicrobiol 2002; 40(6): 1-6.
- Oliviera JE, Silva CA, Oliviera GM, Zaneta DM et al. AntimicrobAgents. Chemoter 2009; 53(7): 2887-289.
- Livermore MD. Pneumonia-causing pathogens and their resistance. Curetis Symposium, ECCMID, 01/04/2014, London, UK.
- RaduPopescu AM, Dumitriu S, Simona ES, Bancescu G, Udristioiu A, Cojocaru M, Vagu C. Phenotypic and GenotipcCaracterisationof Antibiotics. Farmacia2010; (57); 3: 420-427
- RaduPopescuAM, Dumitriu S, Simona ES, Bancescu G, Udristioiu A, Cojocaru M, Vagu C. Resistance PatternsinAcinetobacterbaumani. Strains isolatedin a Romanian Hospital. Farmacia 2010; (58); 4: 422-429
- David A. Relman. Actionable Sequence Data on Infectious Diseases in the Clinical Workplace. ClinChem 2014; 61:38-40.
- Gavin R. Oliver, Steven N. Hart, and Eric W. Klee. Bioinformatics for Clinical Next Generation Sequencing. ClinChem 2014; 61:124-135.



